Research related to induced pluripotent stem cells (iPSCs) is proceeding at warp speed. I have posted multiple blog entries related to iPSCs, ten so far in 2010. These are all listed in the November 2010 blog post Past blog postings on stem cells and epigenomics. In a recent series of comments however(ref), my reader B has made it clear to me that there were still major gaps in my coverage of important iPSC developments over the last year. I fill-in on several of these developments here, occasionally also mentioning research already covered.
Key Developments
Taken together, a series of publications seem to establish:
· Initial passages (test-tube generations) of iPSCs tend to have certain epigenomic differences from corresponding hESCs (human embryonic stem cells) depending on their donor tissue of origin and exhibit a tendency to differentiate back into those donor tissues.
· Genome-wide transcriptional profiles of ESCs and iPSCs can reveal the disparities between them.
· Assuring full pluripotency and equivalence of iPSCs to hESCs can be accomplished by multiple passages or by use of hDAC inhibitors. Thus, there may be a genomic and functional difference between freshly created iPSCs and ones subject to subsequent treatment to assure full pluripotency.
· While there are molecular differences between various strains of iPSCs and ESCs and possible functional differences, some iPSCs are sufficiently pluripotent to create fully functional mice.
· At least some lines of iPSCs exhibit signs of epigenetic youth and significantly delayed senescence. Tissues created from them are correspondingly young. However, several reports of what iPSC reversion does to telomere lengths appear to be contradictory.
· There is at least one report that iPS cells display mitochondrial rejuvenation compared to their fibroblast source cells. This mitochondrial rejuvenation was passed on to fibroblast cells derived from the iPSCs.
· Researchers have succeeded reverted a large number of cell types to iPSC status.
· More reliable and efficient means for cell reprogramming that yield more consistent “higher-fidelity” iPSC are being discovered.
· iPSC research might well lead to therapies for HIV.
· The iPSC research field is still young, vibrant and full of surprises. Although new research results keep pouring in, there is much still to be learned.
· The promise for the future is bright.
Here are some of the relevant publications supporting the above points. Paying attention to the dates of some of these publications and what they say illustrates how thought processes about iPSCs have been evolving just in the course of one year.
Epigenomic characteristics of iPSCs compared to fully pluripotent hESCs
The February 2010 publication Persistent donor cell gene expression among human induced pluripotent stem cells contributes to differences with human embryonic stem cells. “– although hiPSCs have been described as “embryonic stem cell-like”, these cells have a distinct gene expression pattern compared to hESCs, making incomplete reprogramming a potential pitfall. It is unclear to what degree the difference in tissue of origin may contribute to these gene expression differences. To answer these important questions, a careful transcriptional profiling analysis is necessary to investigate the exact reprogramming state of hiPSCs, as well as analysis of the impression, if any, of the tissue of origin on the resulting hiPSCs. In this study, we compare the gene profiles of hiPSCs derived from fetal fibroblasts, neonatal fibroblasts, adipose stem cells, and keratinocytes to their corresponding donor cells and hESCs. Our analysis elucidates the overall degree of reprogramming within each hiPSC line, as well as the “distance” between each hiPSC line and its donor cell. We further identify genes that have a similar mode of regulation in hiPSCs and their corresponding donor cells compared to hESCs, allowing us to specify core sets of donor genes that continue to be expressed in each hiPSC line. We report that residual gene expression of the donor cell type contributes significantly to the differences among hiPSCs and hESCs, and adds to the incompleteness in reprogramming. Specifically, our analysis reveals that fetal fibroblast-derived hiPSCs are closer to hESCs, followed by adipose, neonatal fibroblast, and keratinocyte-derived hiPSCs. — Conclusions and outlook: After analyzing, in detail, genome-wide transcriptional profiles of starting cell populations, partially reprogrammed cells and iPSCs and comparing these with ESCs, we conclude that iPSCs and ESCs share a well-defined core pluripotency network. However, some core genes often seem expressed at lower levels in iPSCs. In addition, this network comprises not only the usual pluripotency transcription factors, but also genes not yet described as, but likely to be, involved in pluripotency and/or self-renewal and genes responsible for many other biological processes, such as cell-cell communication and metabolism.”
he May 2010 publication Aberrant silencing of imprinted genes on chromosome 12qF1 in mouse induced pluripotent stem cells. “By comparing genetically identical mouse ES cells and iPSCs, we show here that their overall messenger RNA and microRNA expression patterns are indistinguishable with the exception of a few transcripts encoded within the imprinted Dlk1-Dio3 gene cluster on chromosome 12qF1, which were aberrantly silenced in most of the iPSC clones. Consistent with a developmental role of the Dlk1-Dio3 gene cluster, these iPSC clones contributed poorly to chimaeras and failed to support the development of entirely iPSC-derived animals (‘all-iPSC mice’). In contrast, iPSC clones with normal expression of the Dlk1-Dio3 cluster contributed to high-grade chimaeras and generated viable all-iPSC mice. Notably, treatment of an iPSC clone that had silenced Dlk1-Dio3 with a histone deacetylase inhibitor reactivated the locus and rescued its ability to support full-term development of all-iPSC mice. Thus, the expression state of a single imprinted gene cluster seems to distinguish most murine iPSCs from ES cells and allows for the prospective identification of iPSC clones that have the full development potential of ES cells.”
The July 2010 publication Cell type of origin influences the molecular and functional properties of mouse induced pluripotent stem cells. “Here we show that iPSCs obtained from mouse fibroblasts, hematopoietic and myogenic cells exhibit distinct transcriptional and epigenetic patterns. Moreover, we demonstrate that cellular origin influences the in vitro differentiation potentials of iPSCs into embryoid bodies and different hematopoietic cell types. Notably, continuous passaging of iPSCs largely attenuates these differences. Our results suggest that early-passage iPSCs retain a transient epigenetic memory of their somatic cells of origin, which manifests as differential gene expression and altered differentiation capacity.”–– “iPSCs derived from different somatic cell types retain a transient epigenetic and transcriptional memory of their cell types of origin at early passage, despite acquiring pluripotent gene expression, transgene-independent growth and the ability to contribute to tissues in chimeras. Continuous passaging resolves these differences giving rise to iPSCs that are molecularly and functionally indistinguishable.”
The August 2010 publication Chromatin structure and gene expression programs of human embryonic and induced pluripotent stem cells. “Recent studies have suggested that ESCs and iPSCs represent different pluripotent states with substantially different gene expression profiles. We describe here a comparison of global chromatin structure and gene expression data for a panel of human ESCs and iPSCs. Genome-wide maps of nucleosomes with histone H3K4me3 and H3K27me3 modifications indicate that there is little difference between ESCs and iPSCs with respect to these marks. Gene expression profiles confirm that the transcriptional programs of ESCs and iPSCs show very few consistent differences. Although some variation in chromatin structure and gene expression was observed in these cell lines, these variations did not serve to distinguish ESCs from iPSCs.”
The September 2010 publication Analysis of human and mouse reprogramming of somatic cells to induced pluripotent stem cells. What is in the plate? For one thing, the introduction to this paper provides a concise summary of progress in generating iPSCs up to the date of the publication. “Briefly, cornerstone publications in the reprogramming field have described the following attributes of iPSCs: they can be transmitted to the germ line [2], generated without the oncogenic factor c-Myc [3], [4], obtained from human cells using the same set of factors [5], [6] as well as other factors [7], obtained without permanent genomic manipulation [8], [9], [10], [11], [12], [13], produced from patient cells [14], [15], [16] even with the correction of a genetic disease [17], and more recently, a study demonstrated that iPSCs can give rise to viable mice by tetraploid complementation assays [18], [19], [20]. Similar to ESCs, iPSC lines have been shown to differentiate into derivatives of the three embryonic germ layers. More specifically, studies have demonstrated iPSC’s ability to generate cells of the cardiovascular and hematopoietic lineages [21], [22], insulin-secreting islet like structures [23], functional cardiomyocytes [24], cells of the neural lineages [25], cells of the adipose lineage [26] and retinal cells [27]. Moreover, a number of papers have began to decipher the mechanisms involved in reprogramming [28], [29], [30], [31], [32], [33], [34], [35], [36], a phenomenon that will likely require significant effort in order to be fully understood.”
“There are a large number of reprogramming experiments published so far encompassing genome-wide transcriptional profiling of the cells of origin, the iPSCs and ESCs, which are used as standards of pluripotent cells and allow us to provide here an in-depth analysis of transcriptional profiles of human and mouse cells before and after reprogramming. When compared to ESCs, iPSCs, as expected, share a common pluripotency/self-renewal network. Perhaps more importantly, they also show differences in the expression of some genes(ref).”
Among things stated under Conclusions and Outlook are: “After analyzing, in detail, genome-wide transcriptional profiles of starting cell populations, partially reprogrammed cells and iPSCs and comparing these with ESCs, we conclude that iPSCs and ESCs share a well-defined core pluripotency network. However, some core genes often seem expressed at lower levels in iPSCs. In addition, this network comprises not only the usual pluripotency transcription factors, but also genes not yet described as, but likely to be, involved in pluripotency and/or self-renewal and genes responsible for many other biological processes, such as cell-cell communication and metabolism. — Although we cannot answer the question of whether iPSCs are truly functionally equivalent to ESCs, it seems increasingly obvious that there exists more than one state of pluripotency. This would explain why we can distinguish between ESCs and iPSCs, but also between iPSCs generated with different protocols. As we believe it is important to select the best iPSCs in terms of their differentiation potential, we propose that checking the newly generated iPSC lines for the silencing of a number of genes marked with bivalent domains would assist in preselecting the most promising iPSCs for further studies. Importantly, even though the field of somatic cell reprogramming moves incredible fast and brings us closer every day to getting the “perfect” protocol for iPSC generation in terms of efficiency, a crucial question remains: will we be able to get cells which are safe to use for therapeutical applications? To answer this question, not only will different cell types, ages and origins have to be tested, but also the protocol used for the generation of the iPSCs. Moreover, understanding the path through which somatic cells arrive to a pluripotent state should allow us to evaluate, more accurately, the potential risks inherent in the use of iPSCs in therapy. The propensity of iPSCs to differentiate and not to go wayward after transplantation, judged by the integrity of their genome and epigenome, will need to be evaluated in great detail(ref).”
The September 2010 publication Epigenetic memory in induced pluripotent stem cells relates “Here we observe that low-passage induced pluripotent stem cells (iPSCs) derived by factor-based reprogramming of adult murine tissues harbour residual DNA methylation signatures characteristic of their somatic tissue of origin, which favours their differentiation along lineages related to the donor cell, while restricting alternative cell fates. Such an ‘epigenetic memory’ of the donor tissue could be reset by differentiation and serial reprogramming, or by treatment of iPSCs with chromatin-modifying drugs. In contrast, the differentiation and methylation of nuclear-transfer-derived pluripotent stem cells were more similar to classical embryonic stem cells than were iPSCs. Our data indicate that nuclear transfer is more effective at establishing the ground state of pluripotency than factor-based reprogramming, which can leave an epigenetic memory of the tissue of origin that may influence efforts at directed differentiation for applications in disease modeling or treatment.”
The December 2010 publication Induced pluripotent stem cells: epigenetic memories and practical implications. “It was generally assumed that iPSCs are functionally equivalent to their embryonic stem cell (ESC) counterparts. Recently, a number of research groups have demonstrated that this is not the case, showing that iPSCs retain ‘epigenetic memory’ of the donor tissue from which they were derived and display skewed differentiation potential. This raises the question whether such cells are fit for experimental, diagnostic or therapeutic purpose. A brief survey of the literature illustrates that differences at both epigenetic and transcriptome level are observed between various pluripotent stem cell populations. Interestingly, iPSC populations with perceived ‘anomalies’ can be coaxed to a more ESC-like cellular state either by continuous passaging-which attenuates these epigenetic differences-or treatment with small molecules that target the machinery responsible for remodelling the genome. This suggests that the establishment of an epigenetic status approximating an ESC counterpart is largely a passive process. The mechanisms responsible remain to be established. Meanwhile, other areas of reprogramming are rapidly evolving such as, trans-differentiation of one somatic cell type to another by the forced expression of key transcription factors.”
At least some forms of iPSCs exhibit significantly delayed senescence though reports of what iPSC reversion does to telomere lengths appear to be contradictory.
The October 2010 publication The LARGE principle of cellular reprogramming: lost, acquired and retained gene expression in foreskin and amniotic fluid-derived human iPS cells has a lot to say, including about this topic. “Cellular reprogramming is a means of assigning greater value to primary AFCs (human amniotic fluid cells) by inducing self-renewal and pluripotency and, thus, bypassing senescence. Here, we report the generation and characterization of human amniotic fluid-derived induced pluripotent stem cells (AFiPSCs) and demonstrate their ability to differentiate into the trophoblast lineage after stimulation with BMP2/BMP4. We further carried out comparative transcriptome analyses of primary human AFCs, AFiPSCs, fibroblast-derived iPSCs (FiPSCs) and embryonic stem cells (ESCs). This revealed that the expression of key senescence-associated genes are down-regulated upon the induction of pluripotency in primary AFCs (AFiPSCs). — — Our study aimed at a more detailed molecular characterization of AFiPSCs. To this end, we generated AFiPSCs and demonstrated their ability to differentiate into the extraembryonic trophoblast lineage. This study also highlights the potential of cellular reprogramming to avert replicative senescence observed in bulk primary AFCs.”
Going on: “— Results: Senescence is bypassed by the derivation of AFiPSCs from human AFCs — To investigate the effect of reprogramming on bypassing senescence observed in primary AFC cultures (Figure 1A-II, -III), we analyzed the expression of senescence and telomere-associated genes in young primary AFCs (P6) and senescent AFC (P17) compared to AFiPSC lines (approximately P20). From a list of 116 senescence-associated genes (Table S4) derived from the Gene Ontology database [35], including those described by Vaziri et al. [45], we identified 64 genes as significantly differentially expressed in AFCs at passage 17 compared to the union of all AFiPSC lines (Figure 5). Of these, telomere-associated genes and genes involved in regulating the cell cycle, e.g. MAD2L2, PARP1, RPA3, DKC1, MSH6, CHEK1, PLK1, POU2F1, CDC2, BLM, WRN, DNMT1, DNMT3B, LMNB1, and CDT1, were down-regulated in primary AFCs compared to AFiPSCs and ESCs. In contrast, PIN1, LMNA, GADD45A, CBX6, NOX4, ENG, HIST2H2BE, CDKN2A, CDKN1A, GDF15 and SERPINE1, among others, were up-regulated in primary AFCs compared to AFiPSCs and ESCs. — Cellular reprogramming bypasses senescence of bulk primary AFCsOne of the great advantages of AFiPSCs over their bulk primary counterparts for any desirable application is their acquisition of the ability to propagate indefinitely. The data presented herein suggest, that this phenotypically rejuvenated appearance of AFiPSCs is based on a gene regulatory network, which averts or at least markedly delays the onset of senescence. This is based on the fact that primary AFCs and AFiPSCs and ESCs exhibit opposing expression patterns related to a large number of senescence-associated genes. In particular, we could detect high expression levels of various cell cycle and telomere elongation-associated genes, such as MAD2L2, PARP1, RPA3, DKC1, MSH6, CHEK1, PLK1, POU2F1, CDC2, LMNB1 and CDT1, as well as TERT itself in AFiPSCs in contrast to primary AFCs(ref).”
This same publication concerned itself with a core pluripotency network shared by ESCs and iPSCs. “AFiPSCs, FiPSCs and ESCs share a core self-renewal gene regulatory network driven by OCT4, SOX2 and NANOG. Nevertheless, these cell types are marked by distinct gene expression signatures. For example, expression of the transcription factors, SIX6, EGR2, PKNOX2, HOXD4, HOXD10, DLX5 and RAXL1, known to regulate developmental processes, are retained in AFiPSCs and FiPSCs. Surprisingly, expression of the self-renewal-associated gene PRDM14 or the developmental processes-regulating genes WNT3A and GSC are restricted to ESCs. — This supports the idea that the main function of KLF4 and c-MYC in the process of reprogramming is to accelerate or enhance the efficiency by increasing a balanced cellular proliferation, while in pluripotent cells they seem to be dispensable [61]–[63]. — Among the expressed genes, which are universally acquired during reprogramming processes, independent of the cell source, are key pluripotency-regulating factors, such as POU5F1, SOX2 and NANOG. These establish a core gene regulatory network essential for maintaining self-renewal and pluripotency [46] (ref).”
A December 2009 publication Telomere dynamics in human cells reprogrammed to pluripotency reported : “IPSCs, like ESCs, have been shown in several reports to display increased activity of at least one important enzymatic component of telomere homeostasis – the reverse transcriptase telomerase (TERT) – compared to the activity seen in somatic cell types (i.e. [14], [15]). More recently it was shown that mouse fibroblasts reprogrammed to pluripotency have both TERT activity and elongated telomeres [9]. This group further demonstrated that although one component of the reprogramming cocktail, the oncogene c-myc, had been shown to directly activate telomerase expression in human cells [17], [18], it was not required for telomere elongation in mouse IPSCs. Marion and colleagues further demonstrated that fibroblasts from both young (6 month) and old donor mice (2.3 yr) elongate telomeres to a similar degree following IPSC conversion.” — Together, these results led the authors to justifiably conclude that “most telomere elongation occurs postreprogramming”[9]. — We examined telomere length in human skin fibroblasts from young and old donor subjects, IPSCs derived from these cells, and IPSCs returned to a differentiated phenotype. We found that like animal cells reprogrammed by either somatic cell nuclear transfer or direct reprogramming, human fibroblasts converted to the IPSC phenotype generally displayed significantly elongated telomeres, and after re-differentiation, displayed a loss of telomere length. Like the mouse, this process was observed irrespective of the inclusion of c-myc in the reprogramming cocktail, and occured to approximately the same degree in cells derived from either young and old subjects. Unlike the mouse however, we observed greater heterogeneity between cell lines, both in the magnitude of telomere elongation during IPSC conversion and telomere shortening following redifferentiation. Also unlike the slow and progressive telomere elongation reported in mouse IPSCs, based on the seven human IPSC lines we analyzed, hIPSC telomeres achieved the 14–15 Kb length characteristic of human ESCs as early as P5.”
The March 2010 paper Telomere elongation in induced pluripotent stem cells from dyskeratosis congenita patients reported consistent results. “Here we show that reprogrammed DC cells overcome a critical limitation in telomerase RNA component (TERC) levels to restore telomere maintenance and self-renewal. We discovered that TERC upregulation is a feature of the pluripotent state, that several telomerase components are targeted by pluripotency-associated transcription factors, and that in autosomal dominant DC, transcriptional silencing accompanies a 3′ deletion at the TERC locus. Our results demonstrate that reprogramming restores telomere elongation in DC cells despite genetic lesions affecting telomerase, and show that strategies to increase TERC expression may be therapeutically beneficial in DC patients.”
The May 2010 publication Spontaneous reversal of the developmental aging of normal human cells following transcriptional reprogramming expresses a quite different view of telomere length restoration in iPSCs. “An hES-derived mortal clonal cell strain EN13 was reprogrammed by SOX2, OCT4 and KLF4. The six resulting induced pluripotent stem (iPS) cell lines were surveyed for telomere length, telomerase activity and telomere-related gene expression. In addition, we measured all these parameters in widely-used hES and iPS cell lines and compared the results to those obtained in the six new isogenic iPS cell lines. — We observed variable but relatively long TRF lengths in three widely studied hES cell lines (16.09-21.1 kb) but markedly shorter TRF lengths (6.4-12.6 kb) in five similarly widely studied iPS cell lines. Transcriptome analysis comparing these hES and iPS cell lines showed modest variation in a small subset of genes implicated in telomere length regulation. However, iPS cell lines consistently showed reduced levels of telomerase activity compared with hES cell lines. — CONCLUSION: Prematurely aged (shortened) telomeres appears to be a common feature of iPS cells created by current pluripotency protocols. However, the spontaneous appearance of lines that express sufficient telomerase activity to extend telomere length may allow the reversal of developmental aging in human cells for use in regenerative medicine.”
Previous blog posts have also cited contradictory reports as to whether telomere lengths in iPSCs are disappointingly short(ref) or amply long (ref). My own conjecture is that all these reports are accurate and the differences are due to differences in the iPSC cell lines studied. A take-away message here is that iPSCs are far from all being the same thing.
More reliable and efficient means for cell reprogramming that yield more consistent “higher-fidelity” iPSC are being discovered.
See the discussions of several additional publications in the July 2010 blog entry Induced pluripotent stem cells – developments on the road to big-time utilization and in the October 2010 entry A breakthrough in producing high-fidelity induced pluripotent stem cells.
iPSC cells display mitochondrial rejuvenation
The November 2010 publication Mitochondrial rejuvenation after induced pluripotency reports “We have examined the properties of mitochondria in two fibroblast lines, corresponding IPSCs, and fibroblasts re-derived from IPSCs using biochemical methods and electron microscopy, and found a dramatic improvement in the quality and function of the mitochondrial complement of the re-derived fibroblasts compared to input fibroblasts. This observation likely stems from two aspects of our experimental design: 1) that the input cell lines used were of advanced cellular age and contained an inefficient mitochondrial complement, and 2) the re-derived fibroblasts were produced using an extensive differentiation regimen that may more closely mimic the degree of growth and maturation found in a developing mammal. CONCLUSIONS/SIGNIFICANCE: These results – coupled with earlier data from our laboratory – suggest that IPSC conversion not only resets the “biological clock”, but can also rejuvenate the energetic capacity of derived cells.”
More is constantly being learned about efficiently reprogramming cells to iPSCs
The October 2010 publication Generation of iPSCs from mouse fibroblasts with a single gene, Oct4, and small molecules reports “Here, we identify a specific chemical combination, which is sufficient to permit reprogramming from mouse embryonic and adult fibroblasts in the presence of a single transcription factor, Oct4, within 20 days, replacing Sox2, Klf4 and c-Myc. The iPSCs generated using this treatment resembled mouse embryonic stem cells in terms of global gene expression profile, epigenetic status and pluripotency both in vitro and in vivo. We also found that 8 days of Oct4 induction was sufficient to enable Oct4-induced reprogramming in the presence of the small molecules, which suggests that reprogramming was initiated within the first 8 days and was independent of continuous exogenous Oct4 expression.”
Researchers have succeeded in reverting a large number of cell types to iPSC status
I cite only a few 2010 and late 2009 publications as examples: Generation of induced pluripotent stem cells by efficient reprogramming of adult bone marrow cells, Effective generation of iPS cells from CD34+ cord blood cells by inhibition of p53, Reprogramming adult hematopoietic cells, and Conversion of ancestral fibroblasts to induced pluripotent stem cells.
More is being learned relative to directing the differentiation of ESCs and iPSCs into target cell lines.
I cite only an example publication, the January 2010 report SIP1 mediates cell-fate decisions between neuroectoderm and mesendoderm in human pluripotent stem cells. Highlights are “* Smad-interacting protein 1 (SIP1) regulates hESC differentiation. * SIP1 upregulation promotes neuroectodermal differentiation. * SIP1 inhibits mesendodermal and endodermal differentiation. * SMAD2/3 and NANOG/OCT4/SOX2 cooperatively regulate SIP1 expression.” Also, “In turn, Activin-Nodal signaling cooperates with NANOG, OCT4, and SOX2 to control the expression of SIP1 in hESCs, thereby limiting the neuroectoderm-promoting effects of SIP1.” The mesendoderm is embryonic tissue which differentiates into mesoderm and endoderm. The mesoderm is “The middle embryonic germ layer, lying between the ectoderm and the endoderm, from which connective tissue, muscle, bone, and the urogenital and circulatory systems develop(ref).” The endoderm is “The innermost of the three primary germ layers of an animal embryo, developing into the gastrointestinal tract, the lungs, and associated structures. Also called hypoblast(ref).” And the ectoderm is “The outermost of the three primary germ layers of an embryo, from which the epidermis, nervous tissue, and, in vertebrates, sense organs develop(ref).”
iPSC research might well lead to therapies for HIV
The November 2010 publication Generation of HIV-1 Resistant and Functional Macrophages From Hematopoietic Stem Cell–derived Induced Pluripotent Stem Cells relates “By developing iPSCs to treat HIV, there is the potential for generating a continuous supply of therapeutic cells for transplantation into HIV-infected patients. In this study, we have used human hematopoietic stem cells (HSCs) to generate anti-HIV gene expressing iPSCs for HIV gene therapy. HSCs were dedifferentiated into continuously growing iPSC lines with four reprogramming factors and a combination anti-HIV lentiviral vector containing a CCR5 short hairpin RNA (shRNA) and a human/rhesus chimeric TRIM5α gene. Upon directed differentiation of the anti-HIV iPSCs toward the hematopoietic lineage, a robust quantity of colony-forming CD133+ HSCs were obtained. These cells were further differentiated into functional end-stage macrophages which displayed a normal phenotypic profile. Upon viral challenge, the anti-HIV iPSC-derived macrophages exhibited strong protection from HIV-1 infection. Here, we demonstrate the ability of iPSCs to develop into HIV-1 resistant immune cells and highlight the potential use of iPSCs for HIV gene and cellular therapies.” Note that since the original donor cells and the redirected iPSC cells were both in the hematopoietic lineage, some of the possible problems of epigenetic traces were obviated.
The scheduled January 2011 publication Human Pluripotent Stem Cells Produce Natural Killer Cells That Mediate Anti-HIV-1 Activity by Utilizing Diverse Cellular Mechanisms offers an additional hope for the battle against HIV. “Natural killer (NK) cells are a key component of the innate immune system with the ability to kill diverse tumor cells and virus-infected cells. While NK cells have been shown to play an important role in the control of HIV-1 replication, their functional activities are often compromised in HIV-1-infected individuals. We have previously demonstrated the derivation of NK cells from human embryonic stem cells (hESCs) with the ability to potently kill multiple types of tumor cells both in vitro and in vivo. We now demonstrate the derivation of functional NK cells from human induced pluripotent stem cells (iPSCs). More importantly, both hESC- and iPSC-derived NK cells are able to inhibit HIV-1 NL4-3 infection of CEM-GFP cells. Additional studies using HIV-1-infected human primary CD4+ T cells illustrated that hESC- and iPSC-derived NK cells suppress HIV-1 infection by at least three distinct cellular mechanisms: killing of infected targets through direct lysis, antibody-dependent cellular cytotoxicity, and production of chemokines and cytokines. Our results establish the potential to utilize hESC- and iPSC-derived NK cells to better understand anti-HIV-1 immunity and provide a novel cellular immunotherapeutic approach to treat HIV/AIDS.”
Promise for the future is bright
The July 2010 publication Evolution of induced pluripotent stem cell technology summarizes progress and the challenges remaining. “iPSC technology provides unprecedented opportunities in biomedical research and regenerative medicine. However, there remain a great deal to learn about iPSC safety, the reprogramming mechanisms, and better ways to direct a specific reprogramming process. The iPSC field will flourish on its mechanistic studies, iPSC-based disease modeling, and identification of new small molecules that modulate reprogramming.”
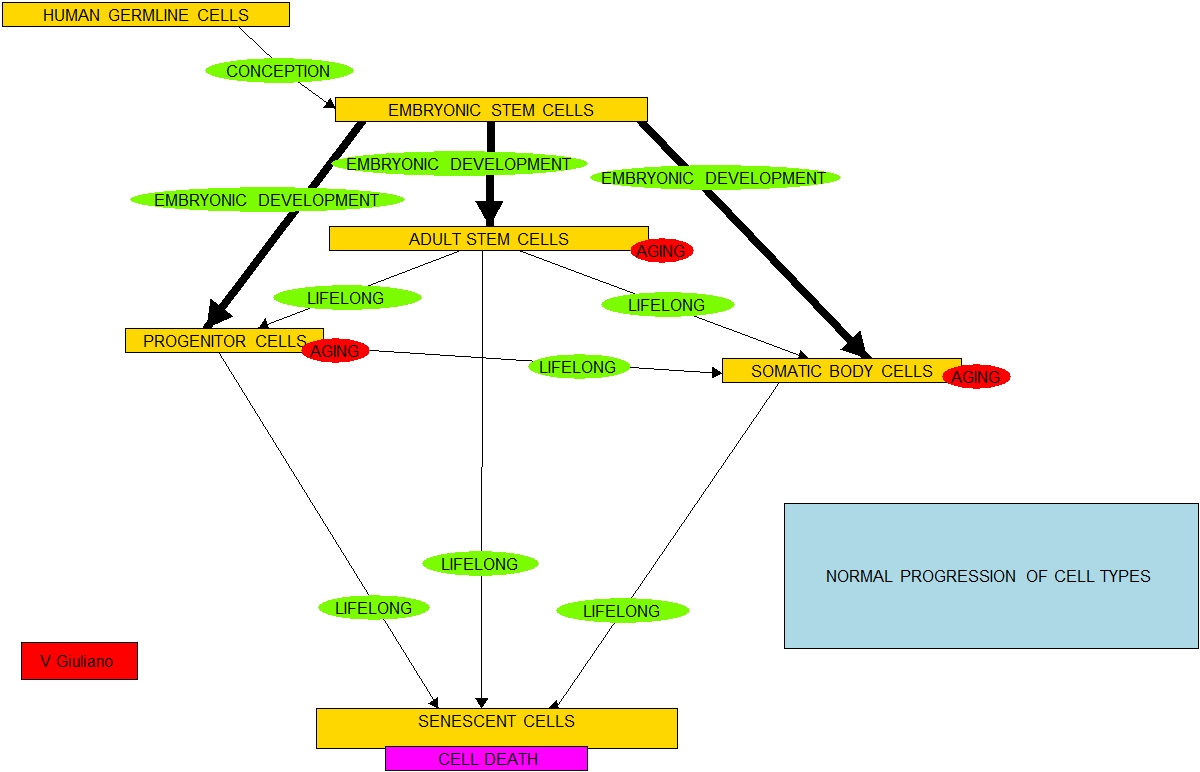
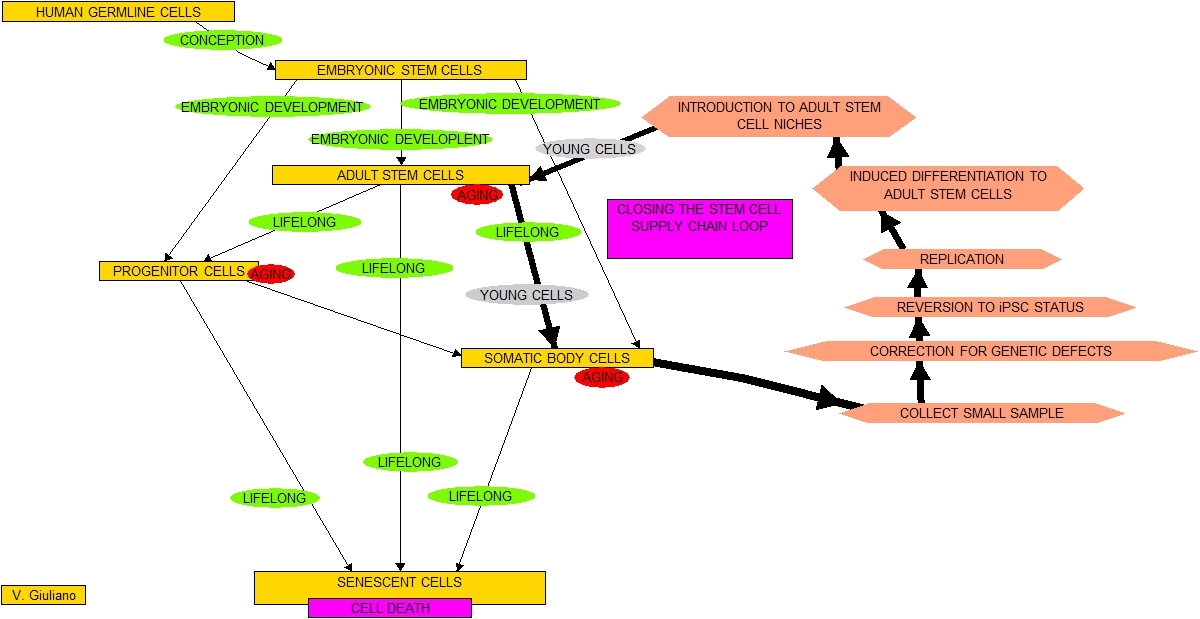
iPSCs have not been ready yet for use in human regenerative medicine experiments although I believe the start of that time is rapidly approaching. I conjecture that If the furious 2010 pace of iPSC research continues we might see animal experiments in the major longevity intervention Closing the loop in the stem cell supply chain in as little as 5-7 years. And the pace appears to be accelerating.