By Vince Giuliano
Recent research indicates a brighter possibility for extraordinary human longevity – healthy lives extending perhaps to hundreds of years. The general approach is one I have been discussing for years, Closing the loop in the stem cell supply chain. Important recent specific progress was brought to my attention by Dr. Victoria Lunyak, a lab director at the Buck Institute who presented on her recent work at the Cell signaling, inflammation and aging symposium in Las Vegas last week, where I was also a presenter. This blog describes key recent work of Dr. Lunyak and others that lends potential reality to the vision of extraordinary human longevity.
Image: The six symbols for longevity
About adult stem cells: relationship to health and longevity and senescence
Life depends on constant replenishment of human body cells with new cells created by differentiation of adult stem cells. Many human body cell types have a remarkable rate of turnover. For example, under normal conditions, red blood cells (RBCs) have a lifespan of about 120 days and from one to five million new red blood cells are produced per second(ref). As Dr.Lunyak pointed out in her presentation, 50-70 million cells are discarded and new ones are generated daily. “In humans, it is estimated that keratinocytes turnover from stem cells to desquamation (skin peeling) every 40–56 days[7] whereas in mice the estimated turnover time is 8–10 days[8](ref).” Health depends critically on the replacement process.
As I originally described the stem cell supply chain:
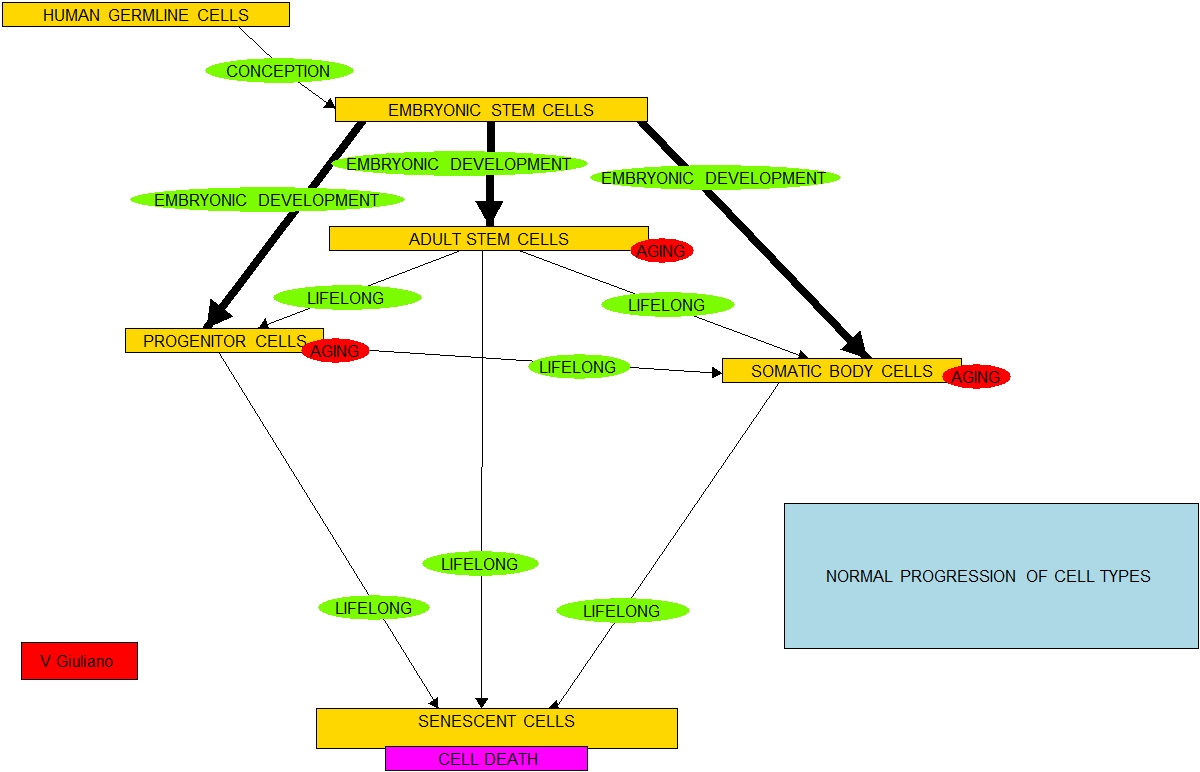
“According to a simplified model of this theory a newly-conceived human embryo consists of pluripotent stem cells (Type A), ones that can potentially divide into any body cells. With growth, these proliferate and, in a remarkably articulated manner, progressively differentiate into multipotent stem cells (Type B), progenitor cells (Type C), mature body somatic cells (Type E), and many eventually become senescent cells (Type E).
According to the best current understanding of stem cells this is an open-loop process. The above list is in order of increasing cell-type specificity and decreasing cell-type potency to differentiate into other cell types. Starting at conception and throughout life, all cells on this list except the senescent ones will selectively reproduce and possibly differentiate into cells of types further down in the list. The state of the body in terms of makeup of cell types continues to change through life and the process goes inexplicably from start (conception) leading to end (death).
At conception, the embryo is all Type A cells. At maturity there are relatively very few Type A cells and a mix of Type B, C and D cells, Type B and C cells typically live in protected stem cell niches where they reproduce and, as-needed differentiate to become the normal working body Type D cells. As Type D cells die from trauma or apoptosis they are replaced by new cells resulting from differentiation of Type B and Type C cells. Stem cell gene expression evolves with age. “In newborn mice, blood-forming cells (hematopoietic stem cells, HSCs) rely on a transcription factor known as Sox17 for self-renewal, but adult HSCs rely on a different transcription factor, Bmi-1(ref).” At an advanced age, the pools of Type B and Type C cells become depleted in part because of replicative senescence and the cells remaining in the pools lose their ability to differentiate as necessary to replace Type D cells.
Although in principle stem cells can replicate indefinitely, in fact they age as the organism ages, continuing to change their gene expression. And the gene expression changes in a way that favors protection against cancer over differentiation capability, e.g. expression of p16ink4a increases. Many Type D cells senesce and become Type E cells which make the corresponding organs shrivel and be susceptible to cancers and other disease processes.
In essence, early-on the body sets up pools of stem and progenitor cells to replace lost somatic cells. Cells in those pools replicate and differentiate throughout life. But when these pools become compromised or depleted or the cells in them lose their capability to differentiate – well – to be blunt soon its curtains.”
That is the essence of the Stem Cell Supply Chain Breakdown theory of aging. IF we could find a way to rejuvenate adult stem cells in their niches, then the stem cell supply chain could possibly be transformed from being a once-through-in-life process to a continuing closed-loop process. And then, human life could possibly go on for a very long time.
This view appears to be consistent with that of some but not all researchers. For example, see Adult stem cells: simply a tool for regenerative medicine or an additional piece in the puzzle of human aging? (Tollervey JR, Lunyak V), Dec. 2011, and also some of the recent technical publications cited below.
Stem cell senescence is an epigenetic phenomenon.
Oxidative stress may well be a factor contributing to hASC senescence, and age-related shifts in niche signaling like Notch is also clearly involved(ref). However, I think the fundamental cause of senescence is epigenetic shifts leading to progressive gene silencing. Senescent cells tend to have shorter telomeres, but I believe this is mainly a downstream and not causal effect.
The 2008 review publication Epigenetic regulation of stem cell fate (Lunyak VV, Rosenfeld MG.) outlines the role of epigenetic factors in adult stem cell senescence and aging. “The multipotency of stem cells is reduced over time due to progressive gene silencing. Genes active in earlier progenitors are gradually silenced at developmentally later stages, and subsets of cell type-specific genes are turned on. This progression is the result of selective expression of transcription factors (TFs) in concert with classis ‘corner stones’ of epigenetics – chromatin remodeling and chromatin modifications, DNA methylation of CpG dinucleotides (4,5). As a result of these events compactization of the chromatin, its accessibility and positioning within specialized nuclear domains undergo dynamic changes. For example, it has been shown that heterochromatic markers, such as HP1 proteins, as well as heterochromatic histone modifications change their localization from dispersed and very dynamic in ESC to more concentrated distinct loci during cellular differentiation (6,7). This suggests that differentiation leads to the restructuring of the chromatin accompanied by the change in the global nuclear architecture, thus allowing the pluripotent nature of ESC genome to become more condensed, and, therefore, more transcriptionally restrained with maturation of the heterochromatin.” – “In mammalian cells, both DNA methylation and specific histone modification are involved in chromatin silencing. DNA methylation and histone modification are believed to be interdependent processes. Recent studies suggest that a combination of histone acetylation and DNA demethylation induces neuronal stem cells (NSC) to trans-differentiate into hematopoietic cells (8).”
Stem cell senescence results from aging and demonstrably leads to diseases and aging.
From the 2011 publication Manifestations and mechanisms of stem cell aging (Liu1,2 and Rando): “Aging is accompanied by a decline in the homeostatic and regenerative capacity of all tissues and organs (Kirkwood, 2005; Rando, 2006). With age, wound healing is slower in the skin, hair turns gray or is lost, skeletal muscle mass and strength decrease, the ratio of cellular constituents in the blood is skewed, and there is a decline in neurogenesis (Sharpless and DePinho, 2007). As the homeostatic and regenerative activities of these tissues are attributable to the resident stem cells, these age-related changes are reflections of declines in stem cell function (Bell and Van Zant, 2004; Dorshkind et. al, 2009; Jones and Rando, 2011). Clearly, in terms of organismal aging, the focus on stem cells is most relevant for those tissues in which normal cellular turnover is very high, such as epithelia of the skin and gut, as opposed to tissues, such as the cerebral cortex and the heart, in which cellular turnover in adults is exceedingly low (Rando, 2006). There is also an increasing interest in the therapeutic potential of stem cells to treat age-related degenerative diseases or conditions, further highlighting the importance of understanding the relationship between stem cell function and the properties of aged tissues.”
Epigenetic interventions can reverse cell senescence markers affecting aging.
According to the publication Histone Demethylase UTX-1 Regulates C. elegans Life Span by Targeting the Insulin/IGF-1 Signaling Pathway (Jin et. al.), 2011, “Like stem cell reprogramming, our results suggest that reestablishment of epigenetic marks lost during aging might help reset the developmental age of animal cells.” Regarding that publication, Lunyak and Kennedy had to say in Aged Worms Erase Epigenetic History: “Defining the molecular events that precipitate multisystem decline is an important component of aging research. In this issue, Jin et al., 2011 show that increased expression of the histone demethylase, utx-1, causes genome-wide decreases in histone H327 trimethylation, which includes the insulin/IGF-1 signaling (IIS) pathway that promotes aging.”
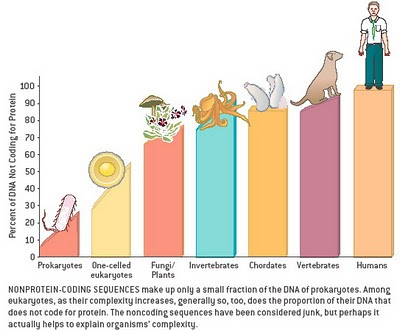
The genome goes far beyond DNA; it includes RNA and all so-called “junk” and non-coding sequences.
As a reminder for us that tend to think only in terms of DNA and genes: “In modern molecular biology and genetics, the genome is the entirety of an organism’s hereditary information. It is encoded either in DNA or, for many types of virus, in RNA. The genome includes both the genes and the non-coding sequences of the DNA/RNA.[1] (ref)”
New categories of both genomic and epigenomic knowledge are emerging, upsetting or refining ideas of just a few years ago.
1. One example is a re-evaluation of the roles of what was once thought to be “junk DNA,” often repeated and transposable segments of DNA not in genes.
The 2011 publication Genomicrelationship between SINE retrotransposons, Pol III-Pol II transcription, and chromatin organization: the journey from junk to jewel (Lunyak VV, Atallah M.) reports: “A typical eukaryotic genome harbors a rich variety of repetitive elements. The most abundant are retrotransposons, mobile retroelements that utilize reverse transcriptase and an RNA intermediate to relocate to a new location within the cellular genomes. A vast majority of the repetitive mammalian genome content has originated from the retrotransposition of SINE (100-300 bp short interspersed nuclear elements that are derived from the structural 7SL RNA or tRNA), LINE (7kb long interspersed nuclear element), and LTR (2-3 kb long terminal repeats) transposable element superfamilies. Broadly labeled as “evolutionary junkyard” or “fossils”, this enigmatic “dark matter” of the genome possesses many yet to be discovered properties.” As described below, such transposable segments may harbor the key to adult stem cell rejuvenation.
Image source
As a matter of fact, human retrotransposons represent for about 13% of the genome and account for about 50% of genetic information, around a million copies (Lunyak PowerPoint presentation). A transposon or “transposable element (TE) is a DNA sequence that can change its relative position (self-transpose) within the genome of a single cell(ref).”
2. Small non-coding RNA species (like IncRNAs, shRNAs, siRNAs and piALU RNAs) can play critical roles in gene regulation, DNA repair and chromatin regulation.
The 2012 publication Protein interactions with piALU RNA indicates putative participation of retroRNA in the cell cycle, DNA repair and chromatin assembly (Blackwell, Luynac et. al.) reports “Recent analyses suggest that transposable element-derived transcripts are processed to yield a variety of small RNA species that play critical functional roles in gene regulation and chromatin organization as well as genome stability and maintenance. Here we report a mass spectrometry analysis of an RNA-affinity complex isolation using a piRNA homologous sequence derived from Alu retrotransposal RNA. Our data point to potential roles for piALU RNAs in DNA repair, cell cycle and chromatin regulations.”
3. In fact, SINE/Alu Retrotransposons are transcriptionally up-regulated upon Senescence of hADSC, and this affects chromatin structure and impairs the DNA damage repair machinery.
(PowerPoint presentation by V. Lunyak). Also the same 2012 publication Protein interactions with piALU RNA indicates putative participation of retroRNA in the cell cycle, DNA repair and chromatin assembly (Blackwell, Lunyak et. al). With hADSC senescence, possibly associated with DNA methylation and age-related chromatin restructuring, there is a loss of regulation controlling SINE/ALU transcription and processing complexes.
Good news covered below is that by knocking down SINE/ALU transcription, hASC senescence can be reversed.
4. Another example is the emerging importance of intragenic DNA methylation, methylation within genes not at promoter sites.
For example, the 2012 publication Intragenic DNA methylation: implications of this epigenetic mechanism for cancer research(Shenker N, Flanagan JM) reports: “Epigenetics is the study of all mechanisms that regulate gene transcription and genome stability that are maintained throughout the cell division, but do not include the DNA sequence itself. The best-studied epigenetic mechanism to date is DNA methylation, where methyl groups are added to the cytosine base within cytosine-guanine dinucleotides (CpG sites). CpGs are frequently clustered in high density (CpG islands (CGIs)) at the promoter of over half of all genes. Current knowledge of transcriptional regulation by DNA methylation centres on its role at the promoter where unmethylated CGIs are present at most actively transcribed genes, whereas hypermethylation of the promoter results in gene repression. — Over the last 5 years, research has gradually incorporated a broader understanding that methylation patterns across the gene (so-called intragenic or gene body methylation) may have a role in transcriptional regulation and efficiency. Numerous genome-wide DNA methylation profiling studies now support this notion, although whether DNA methylation patterns are a cause or consequence of other regulatory mechanisms is not yet clear. This review will examine the evidence for the function of intragenic methylation in gene transcription, and discuss the significance of this in carcinogenesis and for the future use of therapies targeted against DNA methylation.”
Further, the 2012 publication On the presence and role of human gene-body DNA methylation (Jjingo D, Conley AB, Yi SV, Lunyak VV, Jordan IK) reports: “DNA methylation of promoter sequences is a repressive epigenetic mark that down-regulates gene expression. However, DNA methylation is more prevalent within gene-bodies than seen for promoters, and gene-body methylation has been observed to be positively correlated with gene expression levels. This paradox remains unexplained, and accordingly the role of DNA methylation in gene-bodies is poorly understood. We addressed the presence and role of human gene-body DNA methylation using a meta-analysis of human genome-wide methylation, expression and chromatin data sets.”
5. Messenger RNA is subject to adenine methylation and a fifth nucleobases, N6-methyladenosine (m6A), is very common in the transcriptome. Up to 20 percent of human mRNA is routinely methylated. Since more than 5,000 different mRNA molecules contain m6A, mRNA methylation is likely to have widespread effects on how genes are expressed.
The 2012 publication Comprehensive Analysis of mRNA Methylation Reveals Enrichment in 3 UTRs and near Stop Codons (Meyer et. al.) reports “Methylation of the N6 position of adenosine (m6A) is a posttranscriptional modification of RNA with poorly understood prevalence and physiological relevance. The recent discovery that FTO, an obesity risk gene, encodes an m6A demethylase implicates m6A as an important regulator of physiological processes. Here, we present a method for transcriptome-wide m6A localization, which combines m6A-specific methylated RNA immunoprecipitation with next-generation sequencing (MeRIP-Seq). We use this method to identify mRNAs of 7,676 mammalian genes that contain m6A, indicating that m6A is a common base modification of mRNA. The m6A modification exhibits tissue-specific regulation and is markedly increased throughout brain development. We find that m6A sites are enriched near stop codons and in 3 UTRs, and we uncover an association between m6A residues and microRNA-binding sites within 3 UTRs. These findings provide a resource for identifying transcripts that are substrates for adenosine methylation and reveal insights into the epigenetic regulation of the mammalian transcriptome.”
Further, “We found that m6A is present in many mRNAs encoded by genes linked to human diseases, including cancer as well as several brain disorders, such as autism, Alzheimer’s disease, and schizophrenia,” says the study’s lead investigator, Dr. Kate Meyer, a postdoctoral researcher in Dr. Jaffrey’s laboratory(ref).”
6. MicroRNAs can play important roles in tumorigenesis.
The June 2012 publication CpG island hypermethylation-associated silencing of small nucleolar RNAs in human cancer (Ferreira et. al.) reports “Much effort in cancer research has focused on the tiny part of our genome that codes for mRNA. However, it has recently been recognized that microRNAs also contribute decisively to tumorigenesis. Studies have also shown that epigenetic silencing by CpG island hypermethylation of microRNAs with tumor suppressor activities is a common feature of human cancer. The importance of other classes of non-coding RNAs, such as long intergenic ncRNAs (lincRNAs) and transcribed ultraconserved regions (T-UCRs) as altered elements in neoplasia, is also gaining recognition. Thus, we wondered whether there were other ncRNAs undergoing CpG island hypermethylation-associated inactivation in cancer cells”
“Cellular aging is linked to deficiencies in efficient repair of DNA double strand breaks and authentic genome maintenance at the chromatin level. Aging poses a significant threat to adult stem cell function by triggering persistent DNA damage and ultimately cellular senescence.” Further, “65% of naturally occurring repairable DNA damage in self-renewing adult stem cells occurs within transposable elements” of Alu RNA/DNA.
The 2011 publication Inhibition of activated pericentromeric SINE/Alu repeat transcription in senescent human adult stem cells reinstates self-renewal (Lunyak et. al.) reports “Cellular aging is linked to deficiencies in efficient repair of DNA double strand breaks and authentic genome maintenance at the chromatin level. Aging poses a significant threat to adult stem cell function by triggering persistent DNA damage and ultimately cellular senescence. Senescence is often considered to be an irreversible process. Moreover, critical genomic regions engaged in persistent DNA damage accumulation are unknown. Here we report that 65% of naturally occurring repairable DNA damage in self-renewing adult stem cells occurs within transposable elements. Upregulation of Alu retrotransposon transcription upon ex vivo aging causes nuclear cytotoxicity associated with the formation of persistent DNA damage foci and loss of efficient DNA repair in pericentric chromatin. This occurs due to a failure to recruit of condensin I and cohesin complexes. Our results demonstrate that the cytotoxicity of induced Alu repeats is functionally relevant for the human adult stem cell aging. Stable suppression of Alu transcription can reverse the senescent phenotype, reinstating the cells’ self-renewing properties and increasing their plasticity by altering so-called “master” pluripotency regulators.”
According to Wikipedia : “An Alu element is a short stretch of DNA originally characterized by the action of the Alu restrictionendonuclease[1]. Alu elements of different kinds occur in large numbers in primate genomes. In fact, Alu elements are the most abundant Transposable elements in the human genome.”
“Pairs of human chromosomes. 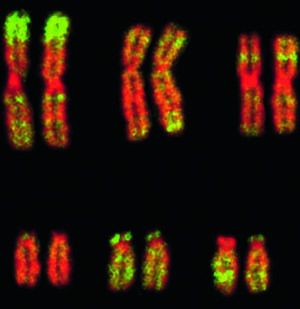 Highlighted in green is alu, a ‘junk’ DNA sequence that distinguishes primates from other mammals.” Image and title source.
Highlighted in green is alu, a ‘junk’ DNA sequence that distinguishes primates from other mammals.” Image and title source.
Specific interventions involving removal of damaged specific segments of RNA, formerly thought to be “junk RNA,” can reverse adult stem cell senescence. Working with specific segments of RNA can add to the traditional epigenetic interventions that mainly have related to DNA methylation and histone acetylation. Specifically, by modifying a Lentivirus genome to express GFP and sh-RNA against Alu transcript, it is possible to knock down the generic SINE/Alu transcript in senescent adult stem cells, reversing senescence markers, rejuvenating the cells, and restoring their lost differentiation capability.
Also from the 2012 publication Protein interactions with piALU RNA indicates putative participation of retroRNA in the cell cycle, DNA repair and chromatin assembly (Blackwell, Luynac et al): “Recently, we have reported that the majority of the repairable DNA damage sites in self-renewing human adult stem cells is associated with the retrotransposal portion of the genome, in particular, Alu retrotransposons.7 Moreover, it has been shown that the upregulation of transcriptional activity from Alu can be triggered by genotoxic stress-induced damage30 as well as upon in-vivo and ex-vivo aging in human retinal pigment epithelium (RPE)31 and human adult stem cells,7 respectively. Intriguingly, evidence indicates that cellular cytotoxicity resulting from increased accumulation of Alu RNA is directly linked to Dicer-1 deficiency,4 suggesting that such cytotoxicity may not necessarily be due to an increase in transcriptional activity, but rather the absence of concomitant transcript processing. Our recently published data indicate that the accumulation of unprocessed Alu transcripts triggers chromatin deterioration, loss of DNA repair in pericentromeric areas eliciting the persistent DNA damage response and, ultimately, cellular senescence.7 — Further, we have shown that human adult stem cells stably expressing an shRNA against Alu transcripts, sh-132Alu, override cellular senescence and reinstate their DNA repair capacity,7 suggesting that RNAi machinery is involved in these events. This published data also implies two equally possible mechanisms through which an shRNA against Alu might mediate the observed function: (1) through the post transcriptional gene silencing (PTGS) Dicer-dependent pathway via the cytoplasmic degradation of Alu RNA or (2) through facilitating transcriptional silencing by recruiting either the Dicer-dependent or Piwidependent arms of the RNAi pathway to act directly on the chromatin as shown in Figure 1A. Both of these pathways are plausible and either could depend on the assembly of single or multiple protein complexes capable of cross-talk with DNAdamage-sensing/repair and centromeric maintenance pathways.”
Graph source: (2011)Inhibition of activated pericentromeric SINE/Alu repeat transcription in senescent human adult stem cells reinstates self-renewal Lunyak et. al. Cell Cycle. 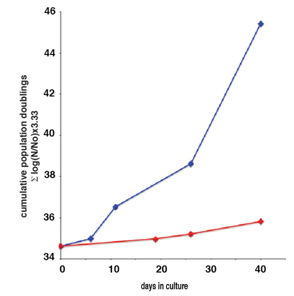 The red line is for senescent hADCs and the blue line shows reinstated proliferation of prior senescent cells.
The red line is for senescent hADCs and the blue line shows reinstated proliferation of prior senescent cells.
Similarly, β-galactoside staining and PCR analyses show that the treated adult stem cells exhibit younger phenotypes.
Wrapping it up
Taking the research all together, it appears that what is happening is adult stem cell rejuvenation via correction of segments of adult stem cell DNA responsible for repair that have themselves become damaged. I think this is a very fundamental line of research. For one matter, it clearly demonstrates epigenetic reversibility of stem cell senescence, a matter some researchers have thought to be impossible. In terms of the Stem Cycle supply chain, the research also suggests that in may not be necessary to revert cells to iPSC status to close the stem cell supply chain loop. It may be sufficient to start with pluripotent adult stem cells, probably adipose-derived ones which are common and relatively easy to collect(ref). These are easier to revert to iPSC status than somatic cells, but such full reversion might not be necessary or desirable.
Of course, further research will be required to develop the in-vivo and eventually human counterparts of this research. How do the formerly senescent adult stem cells expressing an shRNA against Alu transcripts compare with iPSCs? What happens when rejuvinated formerly senescent hADS cells are inserted back into the animal they came from? What will be the consequence on health and longevity? Will there be glitches in the process such as tumorgenesis? Can the entire process of epigenetic reversion of senescent stem cells be safely induced in-vivo? Exactly how? Can this be the basis of an approach to restoring health and youth to humans?
There is still a way to go but the path itself is exciting.


Thanks for the update from this conference.
I think the more approaches the better. The more fronts, more research, more advances on aging. From DNA damage, Cancer, Inflamation, CNS illness, and specific big killers like heart disease the research will increase our lifespan. The question is how much and how fast. Our admitted ignorance can drive us forward with more questions and more insights.
Here is an article of interest.
NF-κB inhibition delays DNA damage–induced senescence and aging in mice
http://www.jci.org/articles/view/45785?key=84c4d84388bea61c718c
Eric 25001
Your comments are right-on Since healthspans and lifespans are tughtly correlated, a wide variety of research publications from main corridors and nooks and crannies of the world’s biomedical and bioscience literature may bear on aging. Put differently, aging is the main risk factor for most degenerative diseases. (Some have claimed for all of such diseases.). So, the research sponsored as “aging research” is only a tiny fraction of that requiring monitorship. A hope of mine for this blog is to intigrate findings from different researchers who don’t know and don’t talk to each other. That is one reason why my blog entries tend to be so long.
As to NF-κB, the publication seems to be very interesting. I have long had an interest in NF-κB and the increase of NF-κB expression with aging and its role in inflammation. This is discussed at some length in my treatise. And I point out there that some 36 substances in my anti-aging firewall supplement regimen are inhibitors of NF-κB. Further, expression of Nrf2 and expression of NF-κB seems to be opposites. Upgrading expression of one tends to inhibit expression of the other. The negative feedback loops between the two are fairly clear. For example, blocking NF-κB upgrades expression of Nrf2 which turns on antioxidant genes, protecting against cell damage. Soon my Las Vegas Powerpoint presentation on Nrf2 should be on-line, and this fact is emphasized in a couple of slides.
Vince
Vince; I am also looking forward to updates on cytosine methylation as it promises to be a marker of aging and might lead to a way or ways to slow, stop, or reverse aging.
As the team reports online today in the Proceedings of the National Academy of Sciences, it found a significantly higher amount of cytosine methylation in the newborn than in the centenarian: 80.5 percent of all cytosine nucleotides, compared with 73 percent. To look at an intermediate case, the team also performed WGBS on the DNA of a 26-year-old male subject; the methylation level was also intermediate, about 78 percent.
Ref.: Holger Heyn et al., Distinct DNA methylomes of newborns and centenarians, Proceedings of the National Academy of Sciences, 2012, DOI: 10.1073/pnas.1120658109 (open access)
interesting read,
http://transcripts.cnn.com/TRANSCRIPTS/0704/22/siu.01.html
Hi Vince,
Nice write up, will the presentations be available.
I like the Buck Institute, a lot of good research seems to come out of there. I check the website once in a while for publications and videos. I saw there was a recent one on stem cells that included Victoria Lunyak, PhD. The work on the eye stem cells is amazing, but going from clinical trials to human use is a few years off.
“The Future Of Regenerative Medicine: What’s Aging Got To Do With It?”
http://vimeo.com/44076894#at=0
I also saw this future conference; would like to go, but budget says otherwise.
Cold Spring Harbor Laboratory
Molecular Genetics of Aging
October 9 – 13, 2012
http://meetings.cshl.edu/meetings/aging12.shtml
JEG3:
The Buck Institute video is indeed fascinating, I quite agree. And as indicated in the post, I think Victoria’s work is of fundamental importance.
As to the Cold Spring Harbor Laboratory conference, yes the program seems fascinating but the cost is considerible, my Summer/Fall schedule is rapidly filling up, and I am giving preference to events that I speak at. I am currently planning to attend the The Ellison Medical Foundation Colloquium on the Biology of Aging August 8-10 in Woods Hole (http://www.ellisonfoundation.org/node/4686) and also have a couple of speaking events in the works which I will announce when firmed up.
Vince.
As to the video for the Las Vegs conference, I still don’t know when it will be available.
Suma
Yes indeed an interesting read, demonstrating how far we have come since and how far we have not come, . It mainly reflects the conventional medical wisdom of the time, 2007, including several still-solid concepts and other statements that I believe are largely discredited now, even though some people hold onto them. In this last category I would put that dietary supplements largely don’t do anything and are a waste of money. Several hundred research studies quoted in many entries in this blog should discredit that though some heavyweight medical people continue to assert it. We have come a long way since then though quacks continue to practice in unregulated countries.
Vince
Vince
Eric25001
The article on cytosine methylation you point to is another chunk of evidence that epigenetic changes in cells are lifelong and part of what appears to be a program of epigenetic evolution that starts with conception and ends with death. What is interesting to me is that the observed overall decrease of DNA methylation, while pronounced, is not terribly drastic (73% for 103 year olds compared to 81% for newborns for CpG methylation). Probably other changes in the epigenome and the details of where the methylation changes are and accumulated DNA damage contribute a lot more to the phenotypes of aging and stem cell senescence. At least, the last point is a suggestion of the Lunyak work.
Vince
Pingback: A stem cell cure for Parkinson’s Disease – so close and yet so far away | AGING SCIENCES – Anti-Aging Firewalls
Pingback: Observations on the evolution of evolution | AGING SCIENCES – Anti-Aging Firewalls
Vince,
I agree with you that the keys to unlocking the complexities of aging may reside in the epigenetics of stem cells. I would add to this the epigenetics of cellular senescence. Here are a few very specific questions that I am trying to find answers to and would enjoy hearing your comments. (You are welcome to continue this dialogue off the blog site, if you wish).
#1 – Differential aging of stem cells in the same niche – HSCs vs MSCs – why?
Why do Hematopoeitic stem cells (HSCs) in the bone marrow mostly maintain their numbers with aging whereas Mesenchymal stem cells (MSCs) in the bone marrow dramatically decline with aging? Obviously the reasons must have something to do with epigenetics and possibly cellular senescence. I have ready that the reason for the age-related decline in BM MSCs is due to a Polycomb Protein-based epigenetic mechanisms. Three Polycomb protein regulated genes: Bmi1, Rae28, and Mel18, appear to regulate self-renewal of specific stem cell types. Polycomb proteins are a “ancient evolutionary epigenetic mechanism” that is important in organisms like D. melanogaster where CpG methylation plays a negligible role in regulating gene expression. If we could “unlock” the cause for this differential decline in stem cells in the BM niche, this may help us unlock the secret to “stem cell decline in function” with aging. Understanding the ancient epigenetic molecular mechanism of Polycomb proteins may be the “key” to understanding differential loss of BM stem cells (i.e. HSCs vs MSCs)
#2 – Resistance to apoptosis by senescent cells – how?
Cellular apoptosis is such an important mechanism of clearing aging cells in our bodies. Without apoptosis occurring, our bodies would be full of old, defective cells. Unfortunately, senescent cells are RESISTANT to apoptosis and will NOT undergo normal programmed cell death. I believe (like I think you do) that the “upstream cause” of cellular senescence is an epigenetic phenomena. In fact the earliest molecular mechanisms that I have heard about with cellular senescence involve the localization of PML-IV/p53/CBP timeric complex and Rb/E2F corepressor complexes to the PML nuclear bodies. How this prevents p53 induced apoptosis is unclear to me. If we could
figure out how to make senescent cells “sensitive” to pro-apoptotic signals (such as p53), then we could “clear senescent cells” and their toxic SASP products.
James Watson:
I have no full answers to your questions but I want to comment that I think they are nonetheless powerful contributions. that is, they create the context for whole new directions of inquiry. Powerful questions in science are sometimes more important than answers which often change with time. What is gravity? Newton had an answer which gave way to Einstein’s more sophisticated answer. Neither answers are satisfactory now given the state of physics new answers of even much more sophistication are emerging.
With regard to both of your questions, answers may be on different levels, including a) what is the evolutionary survival purpose of what is observed? and b) what are the molecular pathway events that create the result? Speaking from evolution with regard to Question #1 I can guess (only a guess) that because hematopoetic stem cells have to do with blood and the immune system, they have to do with immediate short term survival. That is why they turn over so rapidly amd maintain their numbers in their niches with aging. MSCs on the other hand have to do with renewal of bone and other slowly changing tissues and are less concerned with day-to-day survival. So, they are allowed to age and decline in numbers and contribute to overall aging along with numerous other factors. Evolution demands aging and death in virtually all organisms, so this is OK. This is a highly abstract answer and can be questioned. I do agree that looking into the epigenetic mechanisms of the polycomb proteins caould be a good place to start to answer the question in terms of the molecular biology involved.
With regard to Question 2, I can now respond only more speculatively. Perhaps cell senescence is part of an evolutionary program that seems to operate in all species that specifies a maximum lifespans for each species. It is possible that as adult stem cell differentiation slows down with aging, there is associated signaling that inhibits apoptosis in target cells, the idea being that a senescent cell is better than none at all. There is probably something like an epigenetic program for each cell type that operates stochastically but largely determines the epigenetic status of cell types from birth of an organism to death. I am aware of the research that points to how clearing out senescent cells might be life extending. So, as you suggest, insterventions in aged people that upgrade expression of P53 and p21 might be developed that are to some extent life-extending. In essence, we still have very much to learn.
I do look forward to your continuing contributions to this blog.
Vince