By James P Watson with contributions and editing by Vince Giuliano
This blog entry tells two stories. The first story is that of circular RNAs. Once we thought that any important DNA was located in the nucleus of cells on chromosomes, linear strands capped at each end by a telomeres. Of exclusive importance were linear strands of corresponding transcribed RNA for genes which translated into proteins. Now, that view is changing. Over 30 ago, it was observed that a key factor involved in the aging of yeast was the formation of circular fragments of DNA, called extrachromosomal circles r(ERCs), which appeared to serve no function except to exhaust cells of resources. Today we know that there are vast troves of circular non-coding RNAs in human cells, and that they play key roles in epigenetic regulation relating to disease processes and aging. This blog entry outlines this first story. The second story, a far more technical one, is how about one particular piece of circular RNA appears to relate to atherosclerosis and possibly aging – and may be casual of them.
The key messages here are:
- Circular RNA (circRNAs) are abundant, and are found in Human cells – There are between 25,000 and 100,000 circular RNA species per cell! They far outnumber linear RNAs
- CircRNAs are transcribed from DNA but are not translated into proteins
- CircRNAs explains several phenomena observed in DNA: including non-colinear splicing, scrambling of introns, and certain non-coding antisense transcripts.
- CircRNAs may be implicated in disease processes and aging. In particular, splice variants of one long circular RNA known as ANRIL located at the exact location of the 9p21.3 SNP reproduce the same phenotype as the 9p21.3 “risk allele” seen with atherosclerotic disease.
- The research/genetic establishments rejected the idea of circular RNA for a long time, so a great deal is yet to be learned about them.
- CircRNAs are evolutionarily conserved – passed on from generation to generation
- CircRNAs live in to cytoplasm and are long lasting
- CircRNAs offer large number of docking sites for miRNAs, including ones which are capable of silencing genes – they are like coat racks for siRNAs
- The net impact of circRNAs on gene expression can be significant because their siRNA docking sites are competitive with those on genes:
a. Wanted or unwanted gene activation if too many siRNAs of a given kind are docked on circRNAs
b. Wanted or unwanted gene inactivation if too few siRNAs of a given kind are docked on circRNAs or if many siRNAs of a given kind or are released from circRNAs
10. There is emerging evidence that circRNAs tend to function as post-transcriptional regulators of gene expression in specific tissues and during specific developmental phases. This may be due to their miRNA docking capabilities.
11. CircRNAs are particularly expressed in the brain and spinal cord.
12. In yeast, it appears that yeast cell circRNA counterparts accumulate with aging, consume increasing cell resources, and likely cause aging phenotypes. While no such strong causal effect of circRNAs is yet established for mammalian cells, further research into circRNAs and aging is likely to yield interesting results.
PART 1 – THE STORY OF CIRCULAR RNA
Preamble
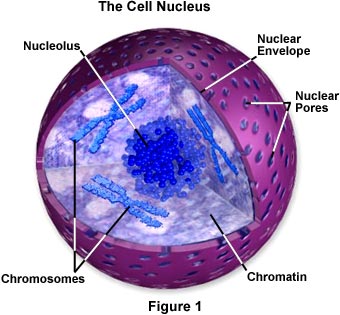
Back in 1993 Leonard Guarante and David Sinclair published Extrachromosomal rDNA circles–a cause of aging in yeast. The abstract is prophetic of what is covered here. “Although many cellular and organismal changes have been described in aging individuals, a precise, molecular cause of aging has yet to be found. A prior study of aging yeast mother cells showed a progressive enlargement and fragmentation of the nucleolus. Here we show that these nucleolar changes are likely due to the accumulation of extrachromosomal rDNA circles (ERCs) in old cells and that, in fact, ERCs cause aging. Mutants for sgs1, the yeast homolog of the Werner’s syndrome gene, accumulate ERCs more rapidly, leading to premature aging and a shorter life span. We speculate on the generality of this molecular cause of aging in higher species, including mammals.”
Cell nucleus showing nucleolus Image soruce
1. Circular RNA are abundant, and are found in Human cells – There are between 25,000 and 100,000 circular RNA species per cell!
Until very recently, no one ever thought that human RNA would make a circle. This did not fit the standard “Watson-Crick” model of DNA-RNA-protein sequence of transcription and then translation. Only viruses and viroids formed circles and these were considered pathogenic. Specifically, circular RNA were once thought to be only associated with RNA viruses – viruses that require no DNA intermediates for their replication cycle (hepatitis C, polio, measles, etc.) or the plant equivalents called “viroids”. Even retroviruses required a DNA intermediate, such as the HIV1 or HIV2 virus. Although there was actually a paper in 1991 that mentioned circular RNA from transcribed genes (Nigro, et al, 1991), very few took it seriously. All that changed in 2013. Recent evidence has shown that human fibroblasts contain more than 25,000 unique circular RNA species that arise from 14.4% of expressed genes! In fact, in many cases there were more circular RNA than their linear RNA copies of the same DNA. In fact, there may be many more than 25,000 species. In the specimens treated with the enzyme RNase R, they detected > 100,000 circular RNAs that were beyond the limits of detection in untreated cells (i.e. more sensitive detection methods will probably bump the number up to > 100,000!).
No one initially believed Dr. Sharpless from U of North Carolina when he tried to publish his manuscript about these human circular RNAs. He had to submit his manuscript to five journals before one would publish his findings. Now there have been at least five pivotal papers on this ” New Multi-Circle Circus” species of RNA. Today, there is no longer any doubt that they exist. Here are some key facts about these “virus-like RNAs”.
references:
- Circular RNAs are abundant, conserved, and associated with ALU repeats “These data show that ecircRNAs are abundant, stable, conserved and nonrandom products of RNA splicing that could be involved in control of gene expression.”
- Assume Nothing: The Tale of Circular RNA “Still, circular RNAs appeared to be exceedingly rare. But Sharpless’ 2012 data implied that circular RNAs were everywhere, representing a substantial fraction of a cell’s transcriptional output. The response from peer reviewers was less than enthusiastic. “I’ve been running my lab now for 12 years,” Sharpless says. “I’ve published in Cell, Nature, NEJM … and I’ve never had reviews like this. They were nasty. They were really mean.”
The basic problem was that circular RNAs were invisible to traditional genomic sequencing methods. In fact, there are many species of circular RNAs:

2. Circular RNAs are evolutionarily conserved and can exceed linear mRNAs of the same species by > 10-fold!
This was what Dr. Sharpless’s original paper said and may be why it was so difficult for him to get the paper published. The technique he used to identify these circular RNAs was high-throughput sequencing of RNAs from ribosome-depleted RNA with and without digestion by RNA exonuclease. He noted that the RNA species that had “non-colinear exons” (i.e. a “back splice”) were enriched by exonuclease digestion of RNA. In vivo (with no exonuclease), these “splice error” RNA species were much more stable than their “no splice error” RNA species. (please see “c” below for an explanation of this). The evidence for the conservation of these circular RNAs came from comparing mice circular RNA loci to those in the human genome. In mice testes, they found 69 circular RNA genes that were exactly orthologous (same location and sequence) to the human cell circular RNA genes. In both species, the circular RNAs were produced from the 2nd exon location for these genes. The enzyme that produces these circular RNAs are the kinases HIPK2 and HIPK3.
References: The same publications:
- Circular RNAs are abundant, conserved, and associated with ALU repeats
- Assume Nothing: The Tale of Circular RNA
3. Circular RNAs are not translated but can be targeted by siRNA specific for circular RNA or by siRNA with common sequences with linear RNAs of the same sequence. However, siRNA involving the ends of linear RNA do not silence circular RNA.
Synthetically created circular RNAs and circular RNAs from RNA viruses can be translated with intact circles, but Dr. Sharpless’s group from UNC did a study to see if endogenously produced circular RNAs translate. Several methods were used to determine the “translatability” of circular RNA. First of all, they examined RNA found in association with ribosomes. Linear RNA were found in association with ribosomes, as expected (circles can go through the protein factory). No circular RNA were found in the ribosomal fraction! Then Dr. Sharpless targeted both linear and circular RNA of the same sequence with various synthetic silencing RNA (siRNA). siRNA that were directed at a unique sequence found only in the linear version did not silence circular RNA. siRNA that targeted the “back splice” region was then used. As expected, this siRNA silenced only the circular RNA. siRNA were then created that bound to a common region found on both the linear and circular RNA. As expected, this silenced both versions.
This adds a whole new complexity to microRNA silencing of gene expression. It is a whole new world out there!
References: The same two publications yet again:
- Circular RNAs are abundant, conserved, and associated with ALU repeats
- Assume Nothing: The Tale of Circular RNA
4. Circular RNAs are predominantly found in the cytoplasm and are very stable.
This was a very interesting finding. The linear and circular versions of four RNAs were compared for stability, after stopping transcription by actinomycin D. The linear transcripts underwent degradation with half lives of < 20 hours, whereas the same RNAs in circular form had half lives that exceeded 48 hours. The circular species also were preferentially located in the cytoplasm, based on FISH studies. This suggests that circular RNA are actively exported out of the nucleus or “spill out” when the nuclear envelope is dissolved during mitosis. All RNA is more stable in the cytoplasm, since there are fewer RNA degrading enzymes there. Thus, it may not be the structure of circular RNA that determines their half life, but their cytoplasmic localization that increases their half life. Whatever the reason, the longer half life of circular RNA enhances its epigenetic impacts as we will see.
5. Circular RNAs are created via a non-canonical mode of RNA splicing.
From Circular RNAs Are the Predominant Transcript Isoform from Hundreds of Human Genes in Diverse Cell Types: “Most human pre-mRNAs are spliced into linear molecules that retain the exon order defined by the genomic sequence. By deep sequencing of RNA from a variety of normal and malignant human cells, we found RNA transcripts from many human genes in which the exons were arranged in a non-canonical order. Statistical estimates and biochemical assays provided strong evidence that a substantial fraction of the spliced transcripts from hundreds of genes are circular RNAs. Our results suggest that a non-canonical mode of RNA splicing, resulting in a circular RNA isoform, is a general feature of the gene expression program in human cells.”

6. Circular RNAs come from large exons; are associated with adjacent introns containing Alu repeats; and are associated with large introns adjacent to the circular RNA exon
Introduction to Alu repeats – the active “jumping genes” in primate evolution
Our genomes are far from stable mainly, it is thought, due to the superposition of DNA copying errors. “A transposable element (TE, transposon or retrotransposon) is a DNA sequence that can change its position within the genome, sometimes creating or reversing mutations and altering the cell’s genome size. Transposition often results in duplication of the TE(ref).”
42-43% of the euchromatin (relatively active chromatin in terms of gene expression) in the human genome is made of repetititve and mobile DNA elements and an additional 18% of the heterochromatin (relatively inactive chromatin in terms of gene expression) in the human genome is made of repetitive and mobile elements. (Compared to only 1.2% of our genome is for protein coding genes). Repetitive DNA refers mainly to “satellite’ and “micro satellite” repeats. Mobile elements refers primarily to transposable elements (TEs). The amount of TEs in the human genome is neither the “world record” for the largest or the “world record” for the smallest amount of repetitive DNA. (For instance, E. coli only has 0.5-1% TEs; whereas corn has 50-80% TEs). Barbara McClintock won the Nobel prize for her work in chromosomal breaks in corn, where she identified the concept of TEs and their role in our genome. Transposable elements are also called “jumping DNA”, because they can move around in the genome. TEs can be further broken down into two major types:
a. DNA transposons – these are no longer active in humans and “can’t jump”. They stopped “jumping” during vertebrate phylogenesis, millions of years ago. For this reason, they are considered “fossil DNA” and are very useful in tracing evolution of speciation. In the future, DNA transposons may be “re-awakened” with “next generation gene therapy”, by using the unique “sleeping beauty” transposon site for specific genetic integration site for gene therapy.
b. Retrotransposons – These are still active in the human genome and are found in both euchromatin and heterochromatin. Retrotransposons make up about 45% of the human genome. To “jump”, they require an RNA polymerase to copy the gene, then the copy can “jump” to a second location (i.e. a copy-and-paste method). There are several subtypes of retrotransposons, including long terminal repeat (LTR) retrotransposons and two subcategories of non-LTR retrotransposons – LINES and SINES. LINES make up 21% of the human genome but most of the half million copies are nonfunctional. Only about 100 copies of the L1 LINE are functional and can “jump” in humans.
SINES make up about 11% of the human genome and there are between 500,000 and 1 million copies of the most common SINE, called an “Alu repeat“. Alu repeats have been dramatically amplified in primate genomes and continue to be amplified at the rate of about 1 insertion every 200 new births. At least 33 germ-line human genetic diseases and 16 cases of cancer have been attributed to this “Alu expansion” and may be responsible for about 0.1-0.3% of human disease, via unequal homologous recombination methods.
Alu repeats found in adjacent introns to circular RNA exons – Complimentary Alu repeats and pairs of Alu repeats more common near circular RNA exons
Dr. Sharpless’s lab found that the introns that flanked the “circularized exons” that the circular RNA were transcribed from had a 2-fold higher chance of having an Alu repeat in it than the “non-circularized exons”. This tendency to be associated with Alu repeats was seen for circular RNA that were made from one exon as well as circular RNAs that were made from multiple exons. In addition, pairs of Alu repeats in the flanking introns were significantly more likely to be complementary (i.e. inverted orientation), vs non complimentary. Circularized exons were 6-fold more likely to contain complimentary Alu repeats compared to control, noncircularized exons.
Large adjacent introns found with circular RNAs; circular RNAs also come from large exons
Another interesting finding from Dr. Sharpless lab was that the introns adjacent to the circular RNA exons tended to be large. On average, they were 3 times as large as introns flanking exons that formed linear RNA transcripts. (p < 10 – 19th power). That is amazing! In addition the exons that formed circular RNA tended to be large, on average being 3 times larger than your average exon size.
My comments: In summary, circular RNAs are most likely to come from large exons flanked by large introns with one or more complimentary Alu repeats.
These features must facilitate circularization. One theory of how this occurs is the concept of an “exon lariat” which “lassos” an exon into a circle. This suggests that circularization occurs due to “exon skipping.”
reference: Circular RNAs are abundant, conserved, and associated with ALU repeats
7. Kinase genes are often circularized – many other genes are also circularized, however
There were many different categories of genes that were circularized into RNAs, but the most common was the protein kinases. Many of these were for Kinase and ATP binding (enriched 5-fold enrichment) or in the category of Protein Kinase C-like (2.3 fold enrichment). However there were ubiquity ligase genes that were also circularized (3 fold enrichment).
Reference: Circular RNAs are abundant, conserved, and associated with ALU repeats
8. Circular RNA explain Non-colinear splicing –“Intron Scrambling” can be explained by circular RNA, and are not due to “splicing
It has been known for a long time that thousands of genes showed evidence of “scrambling”. Initially, this was thought to be due to splicing errors where the splicing machinery in cancer cells went haywire. Then intron scrambing was discovered in non-cancer cells used in controls of the cancer experiments. This was also seen in RNA-seq data, but was attributed to genomic rearrangements such as tandem duplications, which are very common, especially circular RNA can appear to have it’s exons in the “wrong order”. This is why the sequencing data from these RNAs we discarded as “lab error”. Depending on which exons are included in the circular RNA, it can appear that “3” comes before “2”. This work was done at Stanford (see references).
9. Circular RNA explain how non-coding antisense transcripts of certain genes are targets of microRNA
There are a few genes in the genome that have no introns. Once example of this is the intronless CDR1 gene, which codes for a “cerebellar degeneration protein”. A particular miRNA called miR-671 was known to regulate this gene, but the way it regulated the gene could not be figured out. Specifically, it appeared to bind to the “antisense strand of DNA” in the gene. Once it was deduced that CDR1 was transcribed into a circular RNA, then the riddle of how miR-671 bound to the circular RNA solved the gene regulation puzzle. This work was done by Jorgen Kjems lab in Denmark. Here is an illustration:
reference: miRNA-dependent gene silencing involving Ago2-mediated cleavage of a circular antisense RNA
10. Circular RNAs “have no tail” – i.e. they do not have any polyadenylation, which is a feature of the “tail” of linear mRNA
Another reason that circular RNA were overlooked is because the “have no tail”. Specifically, normal mRNA undergo polyadenylation at their tail. Circular RNA do not have this polyadenylate tail.
References:
· The Expanding Repertoire of Circular RNAs
11. Circular RNAs are “trapped” by TRAP electrophoresis – i.e. they get “caught” in a well of melted agarose gel
To further determine that these RNA were indeed circular, they were placed in melted agarose gell “wells” in preformed electrophoresis gell. Circular RNAs cannot “escape” these wells and there therefore “trapped”. Linear RNA, on the other hand, could migrate out of the “TRAPs” and migrate towards the electrical gradient in the electrophoresis gel. Then the wells were analyzed and noted to be enriched in the circular RNA, compared to the controls. This makes any easy way to “catch the circles”
Reference: miRNA-dependent gene silencing involving Ago2-mediated cleavage of a circular antisense RNA
12. Circular RNAs are highly expressed in the brain and spinal cord
This was true for CDR1 as well as many other circular RNAs.
References:
- miRNA-dependent gene silencing involving Ago2-mediated cleavage of a circular antisense RNA
- Assume Nothing: The Tale of Circular RNA
- Circular RNAs are abundant, conserved, and associated with ALU repeats
13. Circular RNAs are not a “histone regulatory mechanism” or a DNA cytosine methylation mechanism of regulation.
This is a truly unique form of gene regulation that appears to be that of a “microRNA sink”, consistent with the ceRNA theory of gene regulation by competing microRNAs for microRNA response elements (MREs) found on coding mRNA and non-coding RNA such as long non-coding RNA (LncRNA) and transcribed pseudogenes.
References:
- miRNA-dependent gene silencing involving Ago2-mediated cleavage of a circular antisense RNA
- Assume Nothing: The Tale of Circular RNA
- Circular RNAs are abundant, conserved, and associated with ALU repeats
- The Expanding Repertoire of Circular RNAs
14. Circular RNA can act as a “microRNA sponge” and soak up microRNA, allowing for other genes to be expressed – Examples:
a. Expression of the ciRS-7 circular RNA “soaks up” the microRNA miR-7, allowing for gene expression repressed by miR-7 to be expressed.
Image source “ciRS-7 sequesters and inactivates miR-7 but is resistant to miR-7 regulation due to its circular nature, whereas miR-671 cleaves ciRS-7, recovering the function of miR-7. Image courtesy of J. Kjems.”
b. Expression of the circular RNA from the gene Sry “soaks up” miR-138, allowing for gene expression repressed by miR-138 to be expressed.
This is the “ceRNA theory” of long, noncoding RNA (LncRNA) and transcribed pseudogenes. Specifically, Kjems from Denmark showed that the gene CDR1 was a gene with one long exon and no introns. It formed circular RNAs that he called “ciRS-7”. ciRS-7 was a 1500 bp circular RNA that could bind to miR-671, since ciRS-7 had a perfect binding site that matched the antisense promoter site on CDR1. miR-671 would promote the cleavage and degradation of ciRS-7 when it bound to the circular RNA. This was not the only thing going on, however. ciRS-7 had 70 partial binding sites (referred to as microRNA response elements, or MREs) that could bind to another miRNA called miR-7. Thus, one circular RNA can “soak up” 70 copies of miR-7 in the cell, allowing for genes that were repressed by miR-7 to be expressed. In this aspect, circular RNA serves as an “amplifier of a repressor eliminator”
A version of the same story occurs with the gene Sry. Sry has been known for some time to have a circular RNA produced from this gene. Sry contains 16 binding sites for miR-138. As a result, when Sry is expressed, one circular RNA can “soak up” as many as 16 copies of microRNA, allowing for gene expression of those genes repressed by miR-138 to be expressed. Again, this is an “amplifier of a repressor eliminator.”
References
- Assume Nothing: The Tale of Circular RNA
- miRNA-dependent gene silencing involving Ago2-mediated cleavage of a circular antisense RNA
- Soaking up the Competition – Circular RNAs as Molecular Sponges for miRNAs
15. Circular RNAs compete with other forms of RNA as decoys for miRNAs. This point is illustrated in this diagram:

Image source “Constraints on evolutionary change in microRNAs. MicroRNAs (miRNAs) lie in a fitness valley constrained by their numerous interactions, which include those with the hairpin structure of the precursor miRNA (pre-miRNA), the many target mRNAs and other RNAs that terminate or modulate miRNA binding to target sequences by competing against them. The latter category includes competing endogenous RNAs (ceRNAs), pseudogene decoys and miRNA mimics.”
16. circRNAs appear to have significant regulatory functions
Consistent with their enormous miRNA binding appetites, circRNAs appear to exercise several tissue and developmental stage-specific regulatory functions, The 2013 publication Circular RNAs are a large class of animal RNAs with regulatory potency reports: : “Circular RNAs (circRNAs) in animals are an enigmatic class of RNA with unknown function. To explore circRNAs systematically, we sequenced and computationally analysed human, mouse and nematode RNA. We detected thousands of well-expressed, stable circRNAs, often showing tissue/developmental-stage-specific expression. Sequence analysis indicated important regulatory functions for circRNAs. We found that a human circRNA, antisense to the cerebellar degeneration-related protein 1 transcript (CDR1as), is densely bound by microRNA (miRNA) effector complexes and harbours 63 conserved binding sites for the ancient miRNA miR-7. Further analyses indicated that CDR1as functions to bind miR-7 in neuronal tissues. Human CDR1as expression in zebrafish impaired midbrain development, similar to knocking down miR-7, suggesting that CDR1as is a miRNA antagonist with a miRNA-binding capacity ten times higher than any other known transcript. Together, our data provide evidence that circRNAs form a large class of post-transcriptional regulators. Numerous circRNAs form by head-to-tail splicing of exons, suggesting previously unrecognized regulatory potential of coding sequences.”
17. Pathogenetic Circular RNA have been found in Plants – Viroids, Viroid-like satellite RNA, and circular chloroplast RNA genes
Circular RNAs have been known to exist for a long time in plants, but were always discarded as pathogens. These circular RNAs were called “Viroids”, as opposed to “viruses” since the viroids were not encapsudated like viruses. Like viruses, viroids replicated autonomously in suceptible plants like RNA viruses in humans (i.e. did not involve DNA), but unlike viruses, viroids did not function as mRNAs. Therefore, viroids would “hijack” the enzymes already present in the plant cells, whereas with viruses, they would synthesize their replication enzymes with genes found in the viruses.
A second form of circular RNA was found in plants called “viroid-like satellite RNA” that were encapsidated and could replicate only with the assistance of a “helper virus”. Here are some features of viroids, viroid-like satellite RNA, and the helper viruses that were required for the viroid-like satellite RNA:
RNA species # of Nucleotides (kb) Structure Replication method
- Viroids 246-375 kb circular RNA no DNA intermediate, rolling-circle type of replication
- Viroid-like satellite RNA 350 kb circular RNA no DNA intermediate, rolling-circule type of replication
- Helper virus 4,500 kb linear RNA non-rolling circle type of replication
Many articles have suggested that these viroid-like satellite RNAs and the viroids are “relics of pre cellular evolution”. If so, then circular RNA may be another example of molecular mechanisms that recapitulate ontogeny.
Later came along the discovery that small RNAS of 18-25 nucleotides in length accumulated in plant tissues that would repress gene expression via RNA silencing. These plant RNAs are now called microRNAs and are analogous to the microRNAs found in animal cells. They are produced in response to plant stress, especially in response to plant microbes.
Although circular RNA not associated with plant infections by viroids have yet to be found, it is known that miRNA exist in plants and that plants can manufacture circular RNA for their viroid invaders. On this basis we suspect that it is only a matter of time before non-pathogenic circular RNA will be found that act as a “microRNA sink” in plants, just like in animals.
References:
- Circular RNAs: Relics of precellular evolution?
- Roles of Plant Small RNAs in Biotic Stress Responses
- RNA silencing in plants
18. RNAs with non-colinear exon sequences associated with repetitive DNA had been found in C elegans earlier, but the authors did not know that the RNA that they were describing was circular RNA
I went back to look at the literature and found the following article:
reference: mRNA Surveillance of Expressed Pseudogenes in C. elegans
Here they describe an “alternatively spliced RNA” from “unproductive gene arrangements” , cytoplasmic pre-mRNAs, and endogenous retroviral and transposon RNAs. These RNAs had non-colinearly arranged exons with repetitive DNA in it. All of this points to circular RNA, but the paper was published in 2005. No one knew about nonpathogenic circular RNA in C. elegans back then. My hypothesis is that Quinn Mitrovich and Philip Anderson of the U of Wisconsin, Madison campus, had actually identified circular RNA, but did not know it. They labeled these genes as “pseudogenes”, when in reality, they were non-coding DNA that produced circular RNA which acted as a “microRNA sink” to allow for other genes to be expressed.
PART 2 – A CIRCULAR RNA IMPLICATED IN ATHEROSCLEROTIC DISEASE AND POSSIBLY AGING
A Circular RNA form of the Long noncoding RNA called ANRIL is found on chromosome 9. Circular forms of ANRIL are found at the 9p21.3 locus of near 3 tumor suppressors (p16, p15, p14) and are associated with CVDz and cellular senescence.
The 9p21.3 locus and CVDz – A true independent “genetic disease susceptibility factor” for atherosclerotic disease that is NOT due to lifestyle factors
From the 2009 publication Sequence Variants on Chromosome 9p21.3 Confer Risk of Atherosclerotic Stroke: “The chromosome 9p21.3 region represents a major risk locus for atherosclerotic stroke. The effect of this locus on stroke seems to be independent of its relationship to CAD and other stroke risk factors. Our findings support a broad role of the 9p21 region in arterial disease.”
There is a single nucleotide polymorphisms (SNPs) located in a non-coding region of chromosome 9 (9p21.3), located at quite a long distance from near three tumor suppressors – p16INK4a, p15INK4b, and p14ARF. Specifically, it is located ~ 120,000 bp away from these genes. (That is “true long distance signaling”). This SNP is evolutionarily conserved in both humans and mice, but the triad of near-by tumor suppressors is slightly different. (In humans the 3 tumor suppressors are p16, p15, and p14; whereas in mice, the triad is p16, p15, and p19). The SNP is called the “INK4a/ARF locus” or the “CDKN2a/b locus” even though it is ~ 120,000 bp way from the actual tumor suppressor genes. The SNP is associated with atherosclerosis, coronary artery disease, stroke, myocardial infarction, and aortic aneurysm. This 9p21.3 locus is an “independent disease susceptibility risk factor” that is NOT dependent on high blood pressure, smoking, obesity, or lipid levels in blood. In other words, this is a “true genetic biomarker”, not an association.
In addition, this “true genetic susceptibility SNP” is evolutionarily conserved in all mammals studied so far – i.e. a “pan genome” SNP. For this reason, there is a lot of interest in this locus. If we could solve the 9p21.3 locus problem in mice, it might also work in humans. At present, it is unclear exactly what the problem at this locus is, since it is so far away from p16, p15, and p14. The discovery of Long non-coding RNA and now the discovery of circular RNAs may explain all of this “long distance suppression” by the SNP.
The 9p21.3 locus and Cellular Senescence
In addition to being associated with atherosclerotic disease, the p16INK4a is a tumor suppressor that is overexpressed with cellular senescence. In fact, it is such a “hallmark of cellular senescence” that it is often used as a “biomarker” of cellular senescence. Specifically, cells that over-express p16INK4a are normally labeled as “senescent” and will not divide. (It should be noted, however, that p53 must also be over-expressed for cellular senescence to occur.) The Mayo Clinic developed a mouse model of accelerated aging where the mouse was prone to aneuploidy due to a defective mitotic spindle apparatus protein. These mice underwent accelerated aging and developed senescent cells that over expressed p16INK4a. When an reporter gene located downstream from p16INK4a produced a protein that could induce apoptosis when the drug AP20187 was added, all of the senescent cells expressing p16INK4a would die. The occasional “selective destruction of p16INK4a expressing cells” effectively removed the senescent cells and kept the phenotype of the mice looking young. Although the mice did not live longer than control mice (they still developed aneuploidy, but this was a separate reason for accelerated aging), they did not display any of the classic signs of aging seen in mice and humans. Thus the p16INK4a locus appears to be the cause of cellular senescence. Since cellular senescence per se, is not considered a gene, no GWAS studies have been done to look at the 9p21.3 SNP and any association with cellular senescence susceptibility, but this could be easily inferred by the data from both mice and humans.
ANRIL splice variants producing circular RNA and Polycomb protein repression of the 9p21.3 locus
The 9p21.3 SNP has been found in a sequence of non-coding DNA that recently was noted to be a “long non-coding RNA” sequence called the
“Antisense Non-coding RNA in the INK4/ARF Locus”, or ANRIL for short. Other than the role for ARF in the development of the optic vasculature, all of the INK4/ARF proteins are thought to be largely dispensable for mammalian development, but play a very important role in restraining the growth of cancer. Specifically, in those with the risk allele for the 9p21.3 SNP, they show a reduced expression of p16INK4a, p15INK4b, ARF, and the long, non-coding RNA called ANRIL. As a result, the individuals who have the “risk allele” of this SNP display an increase in proliferation of monocytes and endothelial cells that is typical of atherosclerosis.
Here are some of the manifestations of a lack of p16INK4a or ARF:
a. p16INK4a deficiency – Intimal hyperplasia of blood vessels following injury – this is the most common reason why bypass grafts clot off in vascular disease and why cardiac blood vessel stents narrow and thrombose.
b. ARF deficiency – atherosclerotic plaque formation
c. HSC proliferation – Proliferation of HSCs in the bone marrow has been associated with p16INK4a and aging
d. Atherosclerosis – p16INK4a and p15INK4a expression prevents atherogenesis via TGF-beta.
How could a non-coding SNP located ~ 120,000 base pairs away from these genes repress their expression? This is where ANRIL comes into the picture. ANRIL has 19 exons with no identified open reading frame. Although cloning the gene has been difficult, a growing number of alternatively spliced transcripts of ANRIL have been identified. The reason why it is difficult to clone the gene is that many of the exons are repetitive DNA. Specifically, many of the exons consist entirely of LINE, SINE, and Alu elements (exons 7,8,12, and 14). Over a decade ago, DePinho and colleagues demonstrated that the p16INK4a locus is potently repressed by Polycomb group (PcG) complexes. This repression is critical for the persistence of somatic stem cells.
This PcG repression is critical for the proliferation of somatic stem cells. Also, it is critical for the self-renewing of pancreatic beta islet cells involved with insulin secretion. Two groups have independently shown that two PcG complexes, PRC-1 and PRC-2, localize to the INK4/ARF locus and repress its activity by the trimethylation of histone 3 at the K27 position (H3K27Me3).
Sharpless and his lab showed in 2010 that different alternatively spliced RNA copies of ANRIL correlated with the Atherosclerotic genotype, whereas others did not. Specifically, mRNA transcripts where the central exons were “skipped”, correlated with the Atherosclerotic genotype seen in the 9p21.3 genotype (i.e. when exons 4-12 were “skipped”, atherosclerotic phenotype occurred). Interesting, they also uncovered circular ANRIL (cANRIL transcripts) that also displayed the Atherosclerotic genotype seen in the 9p21.3 SNPs. These circular RNAs included only the central exons of the gene (Exons 4-12). Interestingly, these circular ANRIL copies had a non-colinear exon order and did not have a polyadenylate tail, which is consistent with the findings of circular RNAs. Also the only parts of the gene that showed up in TaqMan and RNA-seq were the ends of the gene that were not encoded in the circular RNA (Exons 1-3 and exons 13-19).
Conclusion: There is a clear link between the “9p21.3 locus SNP” and all atherosclerotic diseases, such as coronary artery disease, stroke, aortic aneurysm, myocardial infarction, etc. This “susceptibility SNP does not appear to be mediated through any lifestyle factor, such as obesity, smoking, or hyperlipidemia. Curiously, the “9p21.3 locus” is located at a distance of 120,000 base pairs from the tumor suppressors p16INK4a, p15, and p14. The polycomb protein repressors known to repress these tumor suppressors (PRC-1 and PRC-2) cannot account for this “long distance” repression by the 9p21.3 SNP. Expression of circular RNA splice variants of the ANRIL long, non-coding RNA, located at the exact location of the 9p21.3 SNP reproduce the same phenotype as the 9p21.3 “risk allele” seen with atherosclerotic disease. The circular RNA produced are due to “exon skipping” of the proximal and distal exons and produces a circular RNA with non-colinear exon order and no polyadenylate tail. This is the strongest evidence that “Exon skipping” causes circular RNA to form and that circular RNA can cause disease.
MEDICAL DISCLAIMER
FROM TIME TO TIME, THIS BLOG DISCUSSES DISEASE PROCESSES. THE INTENTION OF THOSE DISCUSSIONS IS TO CONVEY CURRENT RESEARCH FINDINGS AND OPINIONS, NOT TO GIVE MEDICAL ADVICE. THE INFORMATION IN POSTS IN THIS BLOG IS NOT A SUBSTITUTE FOR A LICENSED PHYSICIAN’S MEDICAL ADVICE. IF ANY ADVICE, OPINIONS, OR INSTRUCTIONS HEREIN CONFLICT WITH THAT OF A TREATING LICENSED PHYSICIAN, DEFER TO THE OPINION OF THE PHYSICIAN. THIS INFORMATION IS INTENDED FOR PEOPLE IN GOOD HEALTH. IT IS THE READER’S RESPONSIBILITY TO KNOW HIS OR HER MEDICAL HISTORY AND ENSURE THAT ACTIONS OR SUPPLEMENTS HE OR SHE TAKES DO NOT CREATE AN ADVERSE REACTION





Hi Thank you for describing a truly exciting breakthrough. On a trivial point you refer to a number of diferent snp’s by their approximate location on the various chromosones. I have recently seen snp’s described by a format that starts with rs and is followed by a number — ie rs1234. That format allows me to look up the snp on 23andm and snpedia (for example). Why don’t you use that format?
Thanks for sharing this important work and your excitement.
It would be truly elegant if aging in humans was due to accumulation of circular rna’s as is the case for yeast.
JZh Rose
As to SNP nomenclature, your idea is a good one, and I will see if I can get myself to do that. What has held me back has been being in a rush to get blog entries out.
Yes it would be an elegant if circular RNAs were a key active driver of human aging, and indeed they could be. I suspect they are just one of a number of drivers, however, and probably not the key ones for the way we age now. That is, when the average lifespan increases, so could their importance.
Vince
Hmm!
Lifestyle does not effect? Then this would be a seperate pathway from autophagy? and several other aging related phenomena like intermittent fasting, protein cycling and yet unverified carbon 60? So what do these paths have in common and how are they not the same? If both are pro or anti aging can they be more additive or multiplicitive in aging effect? Can synthetic versions be made?
Eric25001
Good questions. Yes, as far as I can imagine now, proliferating circular RNAs would be independent of autophagy and the other phenomena of aging or anti-aging that you mention. Proliferating circular RNA may induce aging because of cellular stress induced by the junk presence and because of competition for siRNA docking sites as pointed out. But such competition for siRNA docking sites may also in some cases be beneficial. My impression is that we really don’t understand the roles of or impacts of circRNAs on human aging yet. That Chapter is yet to be written.
Vince
Thank you for describing a truly exciting breakthrough. IGXE online store
I don’t understand a lot of the details of the genetics; it is way beyond me. But curiously, in a broader sense, you say that the circular RNAs are involved both with aging and tumors. The c60 fed mice lived twice as long and did NOT die of any tumors. Could there be some kind of connection here? Could c60s somehow negate the effects of the circular RNAs on both longevity and tumors?