|
Introduction
Like in many areas of science, the prevailing dogma is often disrupted by emerging evidence which either shows that the theory was completely wrong or that there were exceptions to the rule. This essay is a perfect example of what could be described as an exception to what has been a fundamental “dogma” of genetics; or it could describe a new paradigm that requires the old view to be at least modified.
- The “prevailing dogma” here is that guanine nucleotides only form base pairs with cytosine nucleotides. These make up some of the rungs on the famous Watson/Crick double helix structure of DNA
- The new paradigm or exception to this rule is that multiple guanine nucleotides also spontaneously form a “tetraplex” with a positively charged cation, such as potassium, forming what is referred to as a “Guanine Qaudruplex”, or “G-quadruplex” for short.
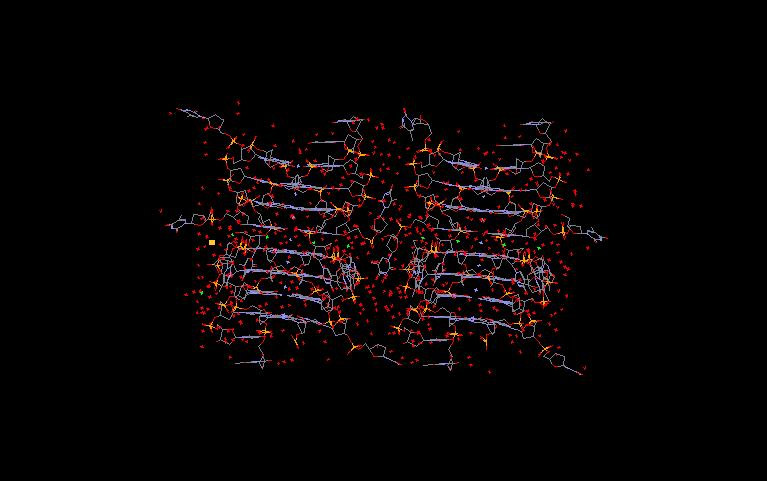
The familiar Double Helix, the Watson-Crick configuration, looks like this:
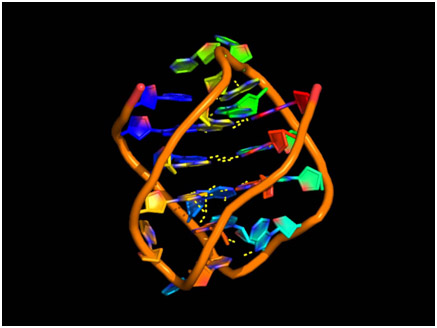
The red-blue rungs are the guanine-cytosine base pairs. Think this is what all or even most DNA looks like? Think again. G-quadraplexes can exist in a variety of topologies that mess up this pretty picture. A G-quadraplex in our DNA could look like these:
So, you can see how considering G-complexes make DNA look a lot more complicated. Actually, the situation is even more complicated since G-quadraplexes are only one of several types of quadraplex structures, albeit the best-understood type. There are also triplex, tetraplex, i-motif, minor groove quadruplexes, etc.
“These unusual DNA structures play critical roles in regulation of very basic biological functions and are integral part of the complex regulatory systems of living beings. The negative supercoiling of DNA can induce sequence-dependent conformational changes that give rise to local DNA structures and alternative DNA conformations such as cruciforms, A-DNA, left-handed DNA (Z-DNA), triplexes, four-stranded DNA (quadruplexes) and others [2,3](ref)” But we don’t really get into most of those other types here.
It is hard to consider G-quadraplex structures as “exceptions to the rule”, since at one or more phases of a mitotic cell’s life cycle, as many as 376,000 of these structures could occur (That is a lot of exceptions to the rule). Moreover, the word “exception” hardly fits here, since these 3-D structures are part of normal molecular biology of both DNA and RNA, and play a key role in the molecular pathogenesis of disease.
Even historically, it has long been known that there was an important alternative to the Watson-Crick model of base pairing, known as Hoogsteen base pairing(ref)(ref) which is the biochemical form of hydrogen bonding involved with G-quadraplexes.
Are G-quadruplexes important? The short answer is YES! They play a key role in telomere stability, telomere length maintenance, hTERT gene expression, oncogene promoter activity, estrogen receptor expression, and many other molecular processes within the cell. In disease, they appear to play key roles in Amyotrophic Lateral Sclerosis, Cancer, and aging. Is that important enough? You bet your G-quadruplexes it is! Here is a recapitulation on what G-quadruplexes are and what they do.
-
G-quadruplexes are an alternative way that DNA and RNA can fold.
The self-association of guanosine bases was first observed in the late 19th century (Ref). In this sense, the spontaneous association of guanosine bases may have been the first “nanotechnology” experiment done. However, the tetrameric arrangement of guanine bases was not determined by X-ray crystallography until 1962, when Gellert, Lipsett, and Davies demonstrated helix formation by guanylic acid (Ref).
The first to show that G-quadruplexes actually occurred in human DNA was Wang and Patel at Columbia University in 1993 (Ref). They showed that the single strand of telomeric DNA that overhangs on the ends of eukaryotic chromosomes form a G-quadruplex. Their study was an artificial DNA solution of the telomere sequence (TTAGGG), formed a “3-stack” of G-quadruplexes at the ends of the telomere (see more about this below under Section 5 on telomeres).
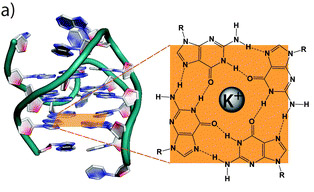
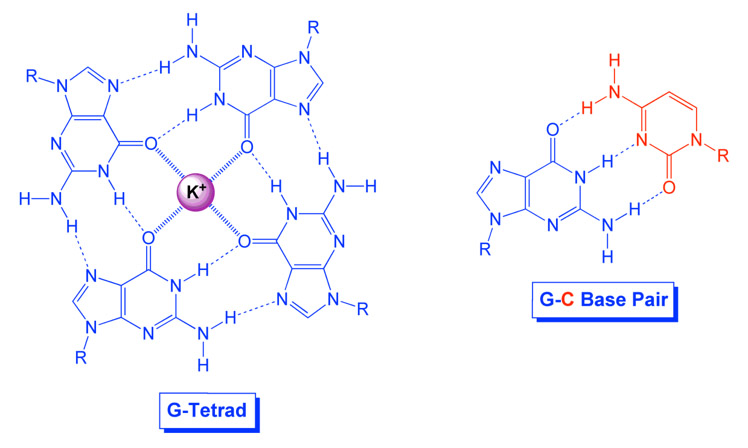
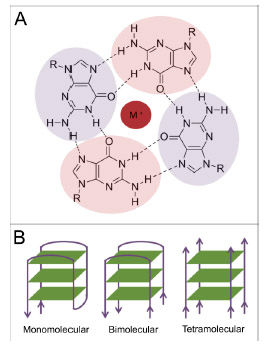
Whereas Guanine normally pairs with cytosine to form one “rung” of the double helix, 4 Guanines can form salt bridges with a metal cation (most commonly potassium) to form a “G-Tetrad”. The bonds between the 4 guanines are called “Hoogsteen base paring“. The cation in the middle and the Hoogsteen base paring are the fundamental features of a a “G-quadruplex”. Here is a diagram comparing a G-C base pair to a G-Tetrad:
Reference: Recognition of Guanine-Rich DNA and RNA by Homologous PNA
-
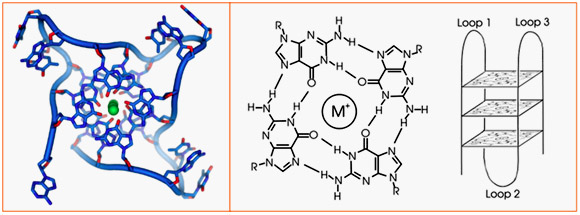
G-quadruplexes are formed from 3 loops of one strand of DNA that is rich in Guanines (Gs).
G-quadruplexes can also form from 2 strands (bimolecular) or 4 strands (tetra molecular) of DNA.
Stabilized by a monovalent (single charged) metal cation, 3 loops of DNA can form a “stack” of G-quadruplexes. The more “stacks” there are in a G-quadruplex, the more stable it becomes. Usually G-quadruplexes with only 1 or 2 stacks are not stable enough to maintain their structure, whereas G-quadruplexes with 3 or more stacks are usually stable. The exceptions to this are the “2-stacks” seen with the thrombin-binding aptamer and fragile X syndrome. Here is an illustration of a stack of 3 G-quadruplexes made from one strand of G-rich DNA:
G-quadraplexes can adopt a variety of geometric configurations depending on where they are found
A good reference on this is the 2007 document Human telomere, oncogenic promoter and 5′-UTR G-quadruplexes: diverse higher order DNA and RNA targets for cancer therapeutics
“DNA can adopt structures other than the Watson–Crick duplex when actively participating in replication, transcription, recombination and damage repair. Of particular interest are guanine-rich regions, which can adopt a non-canonical four-stranded topology called the G-quadruplex. Such architectures are adopted in several key biological contexts, including DNA telomere ends, the purine-rich DNA strands of oncogenic promoter elements, and within RNA 5′-untranslated regions (UTR) in close proximity to translation start sites. Therefore, elucidation of the sequence-based diversity of G-quadruplex scaffolds could provide insights into the distinct biology of guanine-rich sequences within the genome.”
Here is how G-quadruplexes can be formed from 1 strand of DNA, 2 strands of DNA, or 4 strands of DNA:
Reference: Image source DNA and RNA Quadruplex-Binding Proteins
” Four-stranded DNA structures were structurally characterized in vitro by NMR, X-ray and Circular Dichroism spectroscopy in detail. Among the different types of quadruplexes (i-Motifs, minor groove quadruplexes, G-quadruplexes, etc.), the best described are G-quadruplexes which are featured by Hoogsteen base-paring. Sequences with the potential to form quadruplexes are widely present in genome of all organisms. They are found often in repetitive sequences such as telomeric ones, and also in promoter regions and 5′ non-coding sequences. Recently, many proteins with binding affinity to G-quadruplexes have been identified. One of the initially portrayed G-rich regions, the human telomeric sequence (TTAGGG)n, is recognized by many proteins which can modulate telomerase activity. Sequences with the potential to form G-quadruplexes are often located in promoter regions of various oncogenes. The NHE III1 region of the c-MYC promoter has been shown to interact with nucleolin protein as well as other G-quadruplex-binding proteins. A number of G-rich sequences are also present in promoter region of estrogen receptor alpha. In addition to DNA quadruplexes, RNA quadruplexes, which are critical in translational regulation, have also been predicted and observed. For example, the RNA quadruplex formation in telomere-repeat-containing RNA is involved in interaction with TRF2 (telomere repeat binding factor 2) and plays key role in telomere regulation. All these fundamental examples suggest the importance of quadruplex structures in cell processes and their understanding may provide better insight into aging and disease development.”
-
Using prediction computer software, 376,000 putative G-quadruplex DNA structures may be present in the human genome. Common sites include promoters of oncogenes and telomeric DNA. Many non-oncogene promoters also form G-quadruplexes. RNA also forms G-quadruplexes.
Using antibodies that only bind to G-quadruplexes of DNA, these structures have been conclusively shown to exist in human DNA. Here is a fluorescent antibody stain of human osteosarcoma cancer cells which shows the presence of G-quadruplexes:
Image source: 2012 Visualization of DNA G-quadruplex structures in nuclei of human cancer cells
Now since the human genome has been sequenced many times, there are many methods of using software to predict “putative G-quadruplex sequences” (PQS) in the human genome. Using these software tools, Huppert and Balsubramanian at Cambridge University have created a computer program called “quadparser” that uses models of the human DNA sequence.
In 2005, they estimated that as many as 376,000 putative G-quadruplex sequences (PQS) could exist in the human genome. Besides the single stranded DNA overhang of telomeric DNA, the other site where G-quadruplexes show up in the human genome is in promoter regions of oncogenes. For instance, the promoter region for the c-MYC oncogene has G-quadruplexes in it. This may be one of the reasons why c-MYC over-expression is so prevalent in cancer.
Reference: 2005 Prevalence of quadruplexes in the human genome
“Guanine-rich DNA sequences of a particular form have the ability to fold into four-stranded structures called G-quadruplexes. In this paper, we present a working rule to predict which primary sequences can form this structure, and describe a search algorithm to identify such sequences in genomic DNA. We count the number of quadruplexes found in the human genome and compare that with the figure predicted by modelling DNA as a Bernoulli stream or as a Markov chain, using windows of various sizes. We demonstrate that the distribution of loop lengths is significantly different from what would be expected in a random case, providing an indication of the number of potentially relevant quadruplex-forming sequences. In particular, we show that there is a significant repression of quadruplexes in the coding strand of exonic regions, which suggests that quadruplex-forming patterns are disfavoured in sequences that will form RNA.”
-
Many G-quadruplex binding proteins have been found that facilitate G-quadruplex formation or G-quadruplex resolution (de-formation).
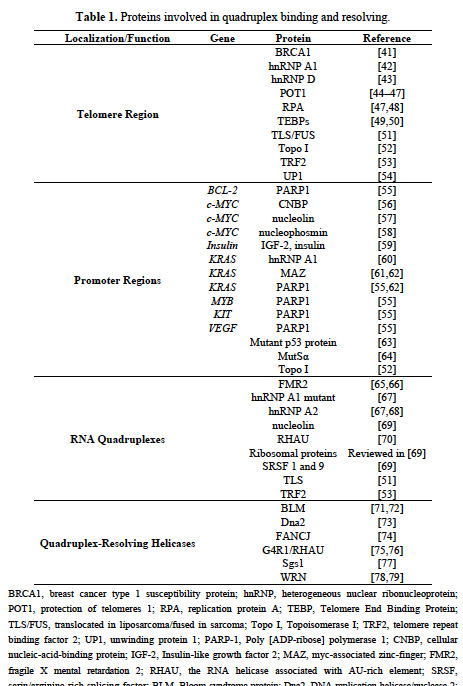
These includes proteins found in the telomere region and promoter regions of DNA, as well as proteins involved with RNA quadruplex formation, and quadruplex-resolving proteins which are called “Helicases“. Here is a list of all of the proteins so far that have been found io be involved with G-quadruplexes in DNA and RNA:
Reference: 2014 DNA and RNA Quadruplex-Binding Proteins
Helicase proteins serve selectively to unwind G-quadraplexes, and when important helicase proteins are missing or mutated, serious diseases can ensue.
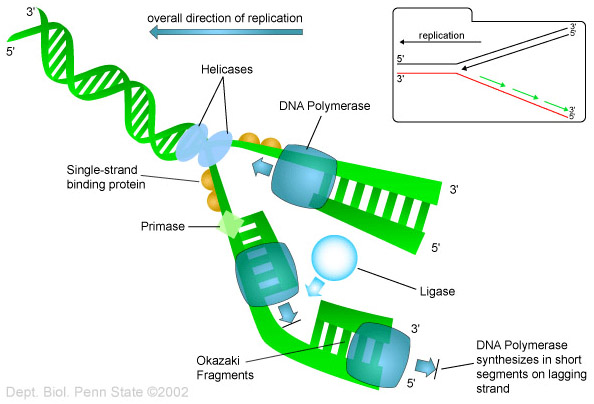
Helicase proteins act as unzippers, to separate intertwined strands of DNA into single strands for such purposes as to allow transcription, replication and recombination. A helicase travels along the DNA in order to unzip it as illustrated here.
G-quadraplexes tend to block many common DNA transactions, again ones such as transcription, replication and recombination. Helicases also unwind G-quadraplexes and often come into play to allow such transactions. Being the major means for doing so, they are discussed throughout this blog entry. For example see Items 8a and 10 below which show how important helicases can be. There is even a mitochondrial “twinkle” helicase for unfolding G-quadraplexes in mitochondrial DNA. See Item 11 below
-
Telomeres are rich in guanines and the single stranded 3′-overhang of DNA forms a DNA G-quadruplex. RNA copies of telomere repeats (i.e. the long noncoding RNA called TERRA) has been shown to form a RNA G-quadruplex.
However, telomeres are not the most common location of G-quadruplexes(82.4%)
5a. Human telomeres contain DNA G-quadruplexes:
Telomeres are made of hexanucleotide repeats with the following sequence in humans: TTAGGG. This means that they are rich in Guanines and therefore form G-quadruplexes. This has been confirmed by many experiments involving specific antibodies and quadruplex binding proteins.
References:
2009 Arrangements of human telomere DNA quadruplex in physiologically relevant K+solutions
2013 Quantitative Visualization of DNA G-quadruplex Structures in Human Cells
“Four-stranded G-quadruplex nucleic acid structures have been of great interest as their high thermodynamic stability under near-physiological conditions suggests that they could form in cells. Here, we report the generation and application of an engineered, structure-specific antibody that was employed to visualize quantitatively DNA G-quadruplex structures in human cells. We explicitly show that G-quadruplex formation in DNA is modulated during cell cycle progression and that endogenous G-quadruplex DNA structures can be stabilized by a small molecule ligand. Together these findings provide substantive evidence for the formation of G-quadruplex structures in the genome of mammalian cells and corroborate the application of stabilizing ligands in a cellular context to target G-quadruplexes and intervene with their function.”
References:
1993 Solution structure of the human telomeric repeat d[AG3(T2AG3)3] G-tetraplex
2009 Selective Recognition of a DNA G-Quadruplex by an Engineered Antibody
2004 Detection of Quadruplex DNA Structures in Human Telomeres by a Fluorescent Carbazole Derivative
2014 Stability of human telomere quadruplexes at high DNA concentrations
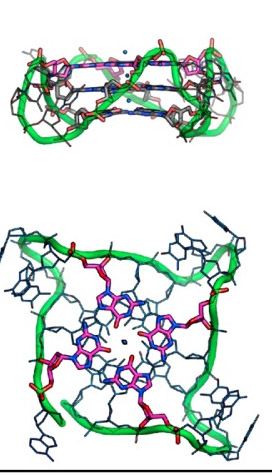
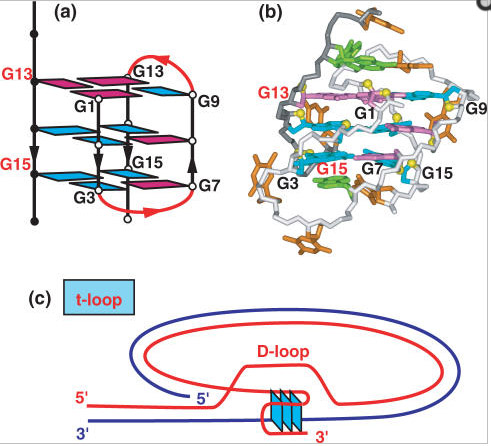
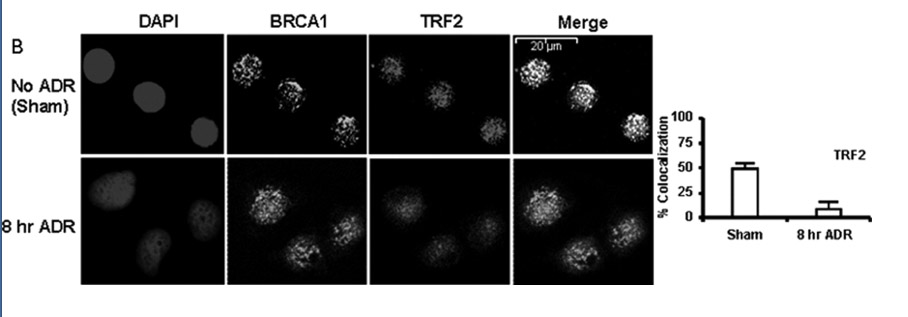
Here is an illustration of the G-quadruplex that forms in human telomeres:
Illustration reference: 2007 Human telomere, oncogenic promoter and 5′-UTR G-quadruplexes: diverse higher order DNA and RNA targets for cancer therapeutics
“Guanine-rich DNA sequences can form G-quadruplexes stabilized by stacked G–G–G–G tetrads in monovalent cation-containing solution. The length and number of individual G-tracts and the length and sequence context of linker residues define the diverse topologies adopted by G-quadruplexes. The review highlights recent solution NMR-based G-quadruplex structures formed by the four-repeat human telomere in K+ solution and the guanine-rich strands of c-myc, c-kit and variant bcl-2 oncogenic promoters, as well as a bimolecular G-quadruplex that targets HIV-1 integrase. Such structure determinations have helped to identify unanticipated scaffolds such as interlocked G-quadruplexes, as well as novel topologies represented by double-chain-reversal and V-shaped loops, triads, mixed tetrads, adenine-mediated pentads and hexads and snap-back G-tetrad alignments. The review also highlights the recent identification of guanine-rich sequences positioned adjacent to translation start sites in 5′-untranslated regions (5′-UTRs) of RNA oncogenic sequences. The activity of the enzyme telomerase, which maintains telomere length, can be negatively regulated through G-quadruplex formation at telomeric ends. The review evaluates progress related to ongoing efforts to identify small molecule drugs that bind and stabilize distinct G-quadruplex scaffolds associated with telomeric and oncogenic sequences, and outlines progress towards identifying recognition principles based on several X-ray-based structures of ligand–G-quadruplex complexes.”
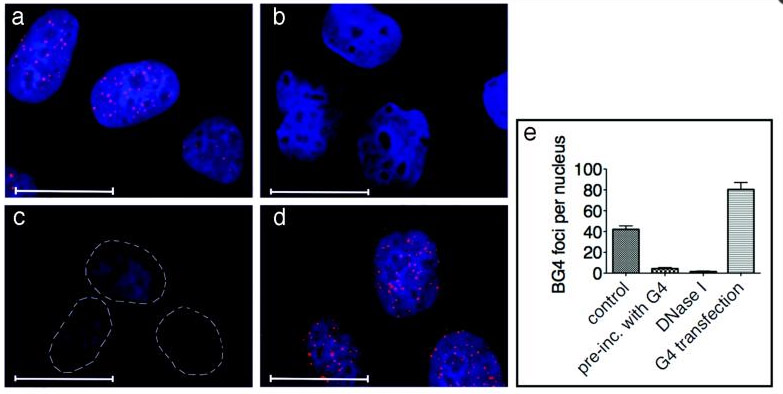
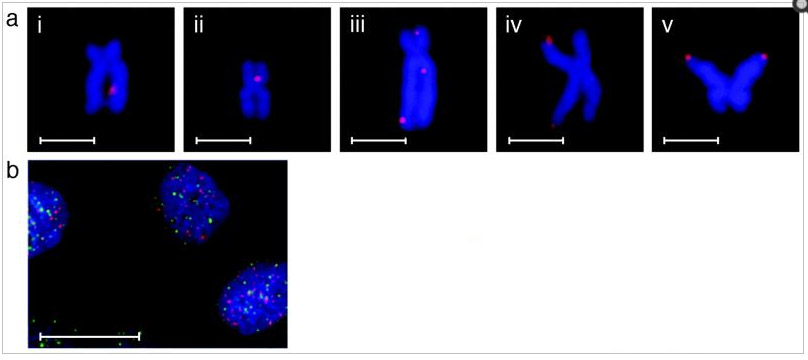
The pictures below are actual visualizations of the G-quadruplexes at telomeres and at non-telomeric regions, as seen in human cancer cells stained with a G-quadruplex antibody. As you can see in iii, iv, and v, there are G-quadruplexes present at telomeres. However, in i, ii, and iii, there are G-quadruplexes present at non-telomeric sites as well. When they quantified the sites, they found that 82.4% of the sites that stained for the G-quadruplexes were not at telomeres. The conclusion of the authors of this article was that most of the G-quadruplexes were found at non-telomeric sites.
Reference: 2013 Quantitative Visualization of DNA G-quadruplex Structures in Human Cells
5b. Telomeric repeats must have at least 3 Guanines to form stable G-quadruplexes (2 are not enough)
Tetrahymena telomeres contain G-quadruplexes but telomeres do not, aat least acording to one publication. This is clearly due to differing G-content The telomeric repeat in Tetrahymena contains 4 guanines (TTGGGG) and forms even more stable G-quadruplexes than human telomeres. However in yeast, the telomeric DNA repeat contains only 2 guanines and it does not form a G-quadruplex.
Reference: 2010 Stability of telomeric G-quadruplexes
“Using biophysical and biochemical methods, we studied sequences mimicking about four repetitions of telomeric motifs from a variety of organisms, including yeasts, with the aim of comparing the G-quadruplex folding potential of telomeric sequences among eukaryotes. G-quadruplex folding did not appear to be a conserved feature among yeast telomeric sequences. By contrast, all known telomeric sequences from eukaryotes other than yeasts folded into G-quadruplexes. Nevertheless, while G3T1-4A repeats (found in a variety of organisms) and G4T2,4 repeats (found in ciliates) folded into stable G-quadruplexes, G-quadruplexes formed by repetitions of G2T2A and G2CT2A motifs (found in many insects and in nematodes, respectively) appeared to be in equilibrium with non-G-quadruplex structures (likely hairpin-duplexes).”
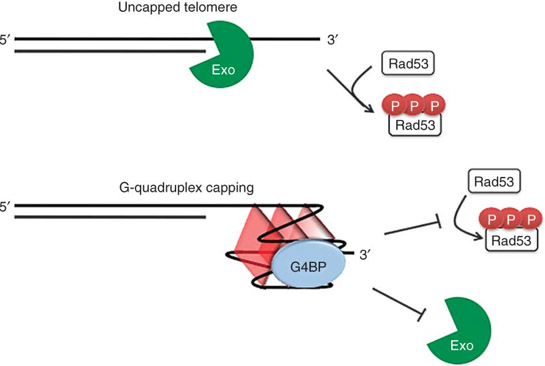
Strangely, the conclusion in this publication that telomeres in yeast do not form G-quadruplexes seems to be contradicted in this 2011 publication which contains an illustration of the telomeric “G-quadruplex capping” that occurs in yeast.
Referemce: 2011 Rudimentary G-quadruplex–based telomere capping in Saccharomyces cerevisiae
” Telomere capping conceals chromosome ends from exonucleases and checkpoints, but the full range of capping mechanisms is not well defined. Telomeres have the potential to form G-quadruplex (G4) DNA, although evidence for telomere G4 DNA function in vivo is limited. In budding yeast, capping requires the Cdc13 protein and is lost at nonpermissive temperatures in cdc13-1 mutants. Here, we use several independent G4 DNA–stabilizing treatments to suppress cdc13-1 capping defects. These include overexpression of three different G4 DNA binding proteins, loss of the G4 DNA unwinding helicase Sgs1, or treatment with small molecule G4 DNA ligands. In vitro, we show that protein-bound G4 DNA at a 3′ overhang inhibits 5′→3′ resection of a paired strand by exonuclease I. These findings demonstrate that, at least in the absence of full natural capping, G4 DNA can play a positive role at telomeres in vivo.”
Also, other publications refer to G-quadraplexes with respect to yeast telomeres like the 2002 publication STM1, a gene which encodes a guanine quadruplex binding protein, interacts with CDC13 in Saccharomyces cerevisiae.
5c. The 3′-overhang of the telomere is where the G-quadruplex forms. TPP1 is a Shelterin protein that helps to form (fold) the G-quadruplex and POT1 is a Shelterin protein that helps to unfold the G-quadruplex.
As it turns out, single strand overhang on the 3′ end of the human telomere contains between 20 and 200 nucleotides. This is where the G-quadruplex forms. Since 3 of the 6 nucleotides in the TTAGGG repeat are guanines, it is easy to see how this could form a G-quadruplex. As it turns out, the G-quadruplex formation of this single stranded DNA is one of the ways that the telomeric DNA is protected from oxidative stress and from triggering the DNA-damage response (DDR), which causes cellular senescence. The formation (folding) of DNA G-quadruplexes and the unfolding of G-quadruplex structures to allow telomerease to work is highly evolved. There are Shelterin proteins that assist in both of these phases of dynamic telomere activity. TRF1 interacting protein 1 (TPP1) assists in the formation of the G-quadruplex and Protection of Telomeres 1 (POT1) protein assists in the unfolding of the G-quadruplex.
References:
2014 DNA and RNA Quadruplex-Binding Proteins
2004 Telomerase Inhibition and Cell Growth Arrest After Telomestatin Treatment in Multiple Myeloma
5d. G-quadraplex formation at the end of telomeric DNA can inhibit telomere extension by telomerase and G-quadraplex unwinding by hellicase.
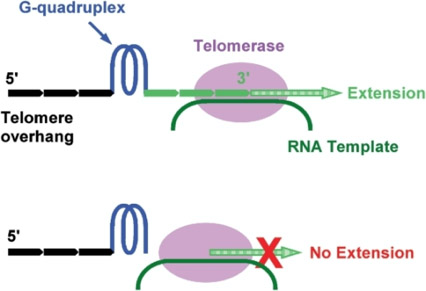
“In this work, we studied the 3′ tail size-dependence of telomere extension by either telomerase or the alternative lengthening of telomere (ALT) mechanism as well as telomere G-quadruplex unwinding. We show that these reactions require a minimal tail of 8, 12 and 6 nt, respectively. Since we have shown that G-quadruplex tends to form at the farthest 3′ distal end of telomere DNA leaving a tail of no more than 5 nt, these results imply that G-quadruplex formation may play a role in regulating reactions at the telomere ends and, as a result, serve as effective drug target for intervening telomere function.”
Image source “Figure 1: Extension of telomere by telomerase depends on the size of single-stranded tail at the 3′ side of the farthest distal G-quadruplex on telomere overhang. A telomere tail of less than four T2AG3 repeats (0–23 nt) will stay in single-stranded form. Those with tails long enough but unable to form G-quadruplex can be extended (top) while others without or with too short tails may not be extended (bottom).”
5e. The BRCA1 protein plays a role in G-quadruplex formation since it regulates the length of the single stranded DNA of the 3′-overhang of the telomere in a Rad50-dependent manner. BRCA1 also controls the expression of the telomerase (hTERT) gene. (i.e. the “telomerease-dependent role of BRCA1).
This is a very important finding, since the BRCA1 gene mutation plays such an important role in inherited breast and ovarian cancer. BRCA1 silencing by epigenetic mechanisms is also seen frequently in sporadic breast cancer as well, making it an important part of the cause of both inherited and sporadic breast cancer pathogenesis. Although the BRCA1 protein plays a key role in repairing double stranded DNA damage, it also inhibits the human TERT promoter region.
BRCA also plays a vital role in controlling telomere length via its association with G-quadruplex formation. These non-DNA repair roles may be why it BRCA1 such a strong influence on the incidence of breast cancer. This is especially intriguing for breast cancer, since the promoter region for the estrogen receptor probably also has G-quadruplexes in it which may also be regulated by BRCA1..
As it turns out, the single strand of the 3′-overhang of the telomere can be as short as 20 nucleotides and as long as 200 nucleotides. Also as it turns out, BRCA1 mutation carriers have longer telomeres and people with normal BRCA1 genes have shorter telomeres. This may be how BRCA1 gene mutations create a “cancer-permissive” environment for cancer to occur.
References:
2009 The Central Region of BRCA1 Binds Preferentially to Supercoiled DNA
2009 BRCA1 Localization to the Telomere and Its Loss from the Telomere in Response to DNA Damage
2003 BRCA1 Inhibition of Telomerase Activity in Cultured Cells
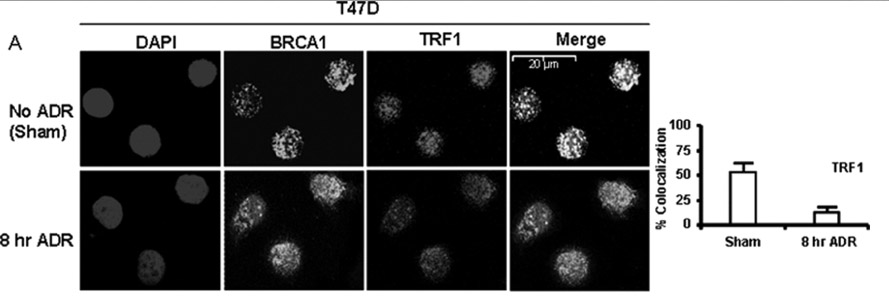
Here are some pictures of the BRCA1 protein co-localizing with TRF1 and with TRF2
Image source: 2009 BRCA1 Localization to the Telomere and Its Loss from the Telomere in Response to DNA Damage
5f. BRCA1 localization at telomeres is lost in response to DNA damage and plays a key role that is “telomerease-independent”
This may be a major mechanism of cancer formation.
Reference: 2009 BRCA1 Localization to the Telomere and Its Loss from the Telomere in Response to DNA Damage
5g. RNA forms G-quadruplexes at telomeres as well (i.e. the lncRNA TERRA)
There is also very strong evidence that the long noncoding RNA transcribed from the antisense strand of telomeres (TERRA) forms RNA G-quadruplexes and that the TERRA RNA G-quadruplex binds to telomere repeat binding factor 2 (TRF2). Here is an illustration of how TERRA forms G-quadruplexes.
-
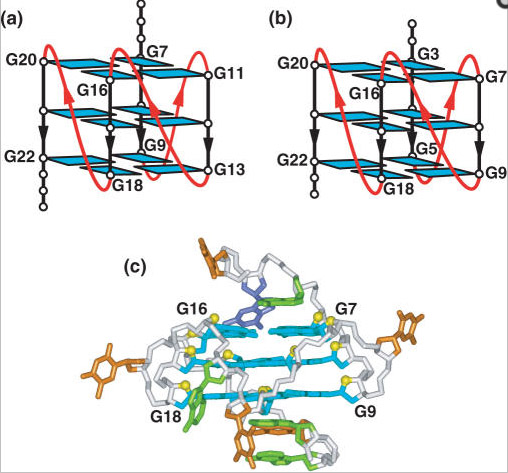
6. The most important oncogene, c-MYC,has a G-quadruplex in it, upstream from the promoter. This 27 nucleotide, G-quadruplex forming region controls 90% of c-Myc gene expression. Thus G-quadruplex formation plays an important role in carcinogenesis.
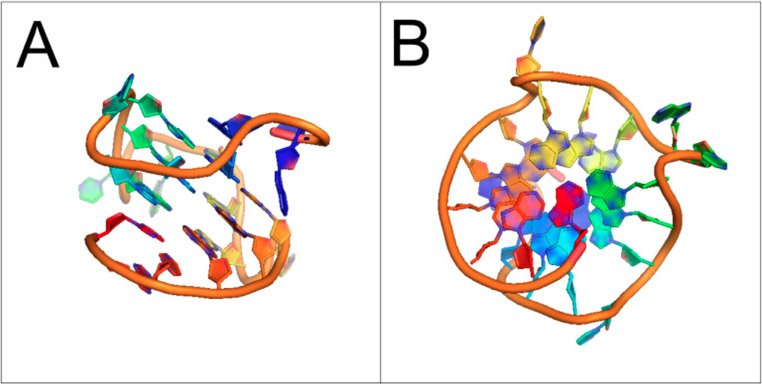
Reference: 2014 DNA and RNA Quadruplex-Binding Proteins “Structure of G-quadruplex in the nuclease hypersensitive element (NHE) III1 region of human c-MYCpromoter (PDBid: 1XAV, [16]). (A) Side view; and (B) Bottom view. Sugar-phosphate backbone is represented by the orange ribbon, with the guanine bases forming the tetrads located in the middle.”
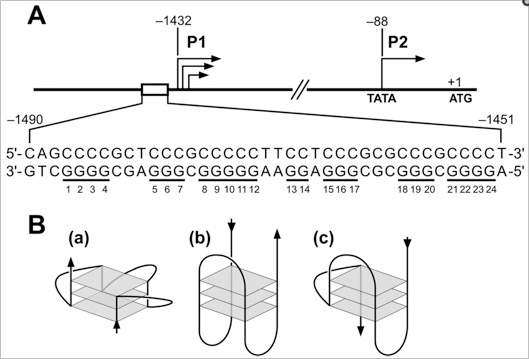
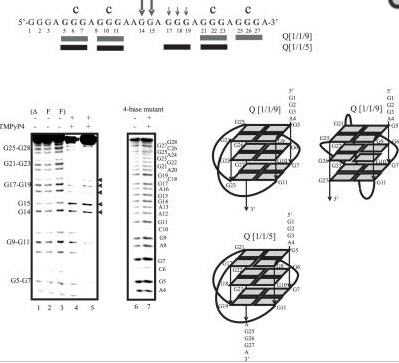
There is a guanine-rich sequence that is upstream from the promoter region of the c-MYC oncogene that forms a G-quadruplex. When this was discovered by Simonsson,Kubista, and Pecinka in Sweden in 1997, suddenly everyone started taking G-quadruplexes seriously!
c-Myc is by far the most important oncogene that is over-expressed in cancer (whereas p53 is the most common tumor suppressor gene that is lost by mutation or by epigenetic silencing in cancer). Moreover, Simonsson and colleagues proposed a molecular mechanism by which an “intrastrand fold-back of DNA” could form a tetraplex with a potassium ion. The exact location where this G-quadruplex can form is located in a major control element, upstream from the promoter, at bases 2186-2212, -115 to -142 bp upstream from the P1 c-Myc promoter. This region is also called the “nuclease-hypersensitive element III” or NHE. Although the c-Myc gene actually has 4 promoters, this NHE accounts for 75-85% of the total c-Myc transcription.
References:
1997 DNA tetraplex formation in the control region of c-myc
Here is an illustration of how the G-quadruplex at the NHE can activate c-Myc gene expression.
Image Reference: 2007 Human telomere, oncogenic promoter and 5′-UTR G-quadruplexes: diverse higher order DNA and RNA targets for cancer therapeutics
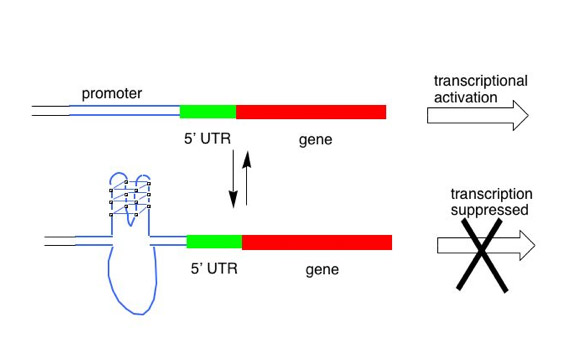
G-quadruplex formation appears to play a role in both gene expression and gene suppression. Here is a Wikipedia illustration how how the G-quadruplex inhibits gene expression. Good examples of genes where G-quadruplex formation inhibits gene expression includes the c-kit gene, the bcd-2 gene, the VEGF gene, and the HIF-1a gene.
Reference: Wikipedia G-Quadraplex
-
Several other common oncogene promoters have G-quadruplex DNA structures upstream from their promoter regions (c-kit, bcl-2, VEGF, HIF-1a, etc.)
c-MYC is not the only oncogene that is regulated by the formation of G-quadruplex DNA structures. Several other oncogenes have well-described G-rich regions that form G-quadruplexes and are involved with gene regulation. Stabilization of G-quadraplexes affecting gene promoter elements to keep them from unwinding is a possible anti-cancer strategy. Here are some examples:
7a. c-kit si an oncogene that codes for a tyrosine kinase receptor with an inhibitory 22 nucleotide G-quadruplex region upstream
The c-kit gene is a common oncogene that codes for the expression of a tyrosine kinase receptor. This gene is often over-expressed in gastrointestinal tumors. The gene has two guanine-rich sequences that can form “3-stack” G-quadruplexes. The formation of the G-quadruplex in the c-kit gene inhibits its expression. Drugs have been developed that stabilize this G-quadruplex structure. These drugs cause the cancer to die by apoptosis. Thus the G-quadruplex of the c-Myc gene and the c-kit gene are structurally different.
References:
2006 A conserved quadruplex motif located in a transcription activation site of the human c-kit oncogene
This publication cites an instance of evolutionary conservation, suggesting similar roles of certain G-quadraplexes in mamalian species: “Mutational analysis of c-kit21 has provided insights into its structural polymorphism. In particular, one mutated form appears to form a single quadruplex species that adopts a parallel conformation. The quadruplex-forming sequence shows a high level of sequence conservation across human, mouse, rat, and chimpanzee. The small variation in sequence between the quadruplex in human/chimpanzee as compared to the rat/mouse was examined more closely by biophysical methods. Despite a variation in the sequence and length of loop 2, the quadruplexes showed both comparable CD spectra, indicative of parallel quadruplexes, and also similar thermal-stability profiles, suggesting conservation of biophysical characteristics. Collectively, the evidence suggests that this quadruplex is a serious target for a detailed functional investigation at the cell-biology level.”
Reference: 2007 Structure of an unprecedented G-quadruplex scaffold in the human c-kit promoter
7b. bcl-2 is an apoptosis inhibitor that is over expressed in cancer and has an inhibitory 39 nucleotide G-quadruplex in it
Cells have many genes that encode for apoptosis activators and apoptosis inhibitors. One of the major apoptosis inhibitor genes is the bcl-2 protein, encoded by the bcl-2 gene. It can act like an oncogene in cancer when it is over expressed. It is now clear that the 39-nucleotide G-quadruplex structure found between 1386 and 1423 base pairs upstream from the transcription start site (TSS) of the bcl-2 gene acts as a bcl-2 gene silencer. When mutation studies have been done that delete this 39-nucleotide sequence, this increases bcl-2 gene expression by 2.1 to 2.6 fold. Like many other genes this guanine-rich sequence is in the proximity of a nuclease hypersensitive region (HNE).
References:
Different G-quadraplexes with alternative configurations can come into play impacting on the bcl-2 gene promoter. “Mutation and deletion analysis permitted isolation and identification of three overlapping DNA sequences within this element that formed the three individual G-quadruplexes. Each of these was characterized using nondenaturing gel analysis, DMS footprinting, and circular dichroism. The central G-quadruplex, which is the most stable, forms a mixed parallel/antiparallel structure consisting of three tetrads connected by loops of one, seven, and three bases. Three different G-quadruplex-interactive agents were found to further stabilize these structures, with individual selectivity toward one or more of these G-quadruplexes. Collectively, these results suggest that the multiple G-quadruplexes identified in the promoter region of the bcl-2 gene are likely to play a similar role to the G-quadruplexes in the c-myc promoter in that their formation could serve to modulate gene transcription. Last, we demonstrate that the complexity of the G-quadruplexes in the bcl-2promoter extends beyond the ability to form any one of three separate G-quadruplexes to each having the capacity to form either three or six different loop isomers. ”
Here is an illustration of a bcl-2 gene G-Quadruplex
Image soure: 2006 Deconvoluting the Structural and Drug-Recognition Complexity of the G-Quadruplex-Forming Region Upstream of the bcl-2 P1 Promoter
7c. The KRAS proto-oncogene has a G-rich region which can activate KRAS gene transcription in the DNA double stranded conformation (i.e. no quadruplex) or inhibit KRAS gene transcription (in the G-quadruplex conformation.
Like the c-kit and bcl-2 G-quadruplexes that regulate gene expression of the c-kit and bcl-2 genes, the KRAS gene also has a guanine-rich strand that can form a “3-stack” G-quadruplex. When this G-quadruplex is stabilized by small molecules such as TMPyP4, it reduces gene expression of the KRAS gene down to 20-40% of controls. When the G-quadruplex is not stabilized, this G-rich area has been shown to go back to the DNA double stranded (Watson-Crick) conformation, thereby allowing gene transcription to occur.
Reference: 2006 G-quadruplex formation within the promoter of the KRAS proto-oncogene and its effect on transcription
Here is a diagram of the 28 nucleotide G-quadruplex that occurs in the KRAS promoter region
Image source: 2006 G-quadruplex formation within the promoter of the KRAS proto-oncogene and its effect on transcription
7d. The VEGF gene and HIF-1a gene each have 5 sites upstream with at least 3 guanines that may form G-quadruplexes, thereby creating or opening up binding sites for Sp1 and Egr-1 transcription factors
One universal feature of cancer is the formation of blood vessels, which provides nutrients and oxygen to the cancer cells, allowing them to grow. A second (near universal feature) of cancer is that 95% of the cancers develop “Warburg-type metabolism”, which is defined as the metabolic phenotype of “aerobic glycolysis in the presence of oxygen”. In other words, they utilized glucose to make ATP without using oxygen, even if there is oxygen present. This metabolic phenomena was named after Otto Warburg who first noted it in the early 20th century. It is now clear that these two “twin features” of cancer (angiogenesis and Warburg-type metabolism) are driven by “twin genes” – VEGF and HIF-1a. For instance, the Vascular endothelial growth factor (VEGF) gene encodes for a protein (VEGF) that is the main transcription factor for making blood vessels. The Hypoxia Inducing Factor 1 alpha (HIF-1a) gene encodes for a protein (HIF-1a) that is the main transcription factor for VEGF and for the many genes that induce the Warburg effect. Interestingly, both of these genes have 5 guanine-rich regions upstream from their promoter that are all capable of forming G-quadruplex structures in vitro. These G-rich regions are near (but not exactly at) endonuclease hypersensitivity regions, suggesting that they are active in their involvement of gene regulation. Here are the DNA sequences of these G-rich regions in VEGF and HIF-1a genes:
VEGF G-rich region(s): The most well-studied G-rich region in the VEGF gene is located at -85 to -50 bps from the TSS
Here is the DNA sequence for this G-rich region:
References:
HIF-1a G-rich regions(s)
Unlike the G-quadruplexes in c-kit and bcl-2 which reduce oncogene expression, these G-rich sequences of the VEGF and HIF-1a genes may actually open up binding sites for promoters that activate these genes, such as Sp-1 and Egr-1 transcription factors.
More references:
“Collectively, our results provide evidence that specific G-quadruplex structures can be formed in the VEGF promoter region, and that the transcription of this gene can be controlled by ligand-mediated G-quadruplex stabilization. Our results also provide further support for the idea that G-quadruplex structures may play structural roles in vivo and therefore might provide insight into novel methodologies for rational drug design.”
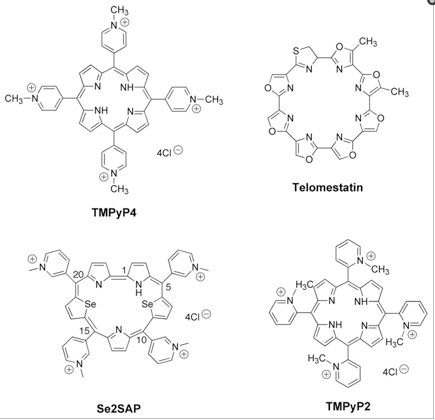
7e. Small molecules have been made that bind to G-quadruplexes of oncogenes
Several small molecules have been synthesized that bind to G-quadruplexes. These molecules actually stabilize the G-quadruplex, which makes them more difficult to “unfold”. As a consequence of this G-quadruplex stability, these small molecules induce apoptosis in cancer cells. It is likely that the location where these work is at the promoters or upstream from the promoters in the oncogenes that have G-quadruplexes in them, such as c-Myc, c-kit, bcl-2, VEGF, and HIF-1a.
References:
Here are some of the small molecules that have been synthesized to bind to G-quadruplexes:
-
G-quadruplexes or G-quadruplex unfolding problems play a key role in many disease conditions (Fragile X syndrome, Werner’s syndrome, Friedrich’s ataxia, ALS, etc.)
Whereas most of the information above involved either telomere G-quadruplexes or cancer oncogenes, several rare neurodegenerative diseases have been found to form G-quadruplexes. In most all of these diseases, a nucleotide repeat expansion occurs where a triplet nulceotide sequence expands or a hexanucleotide repeat expands. The repeat expansions have been found in coding segments (i.e. exons), 5′ untranslated regions (5′-UTRs), 3′ untranslated regions (3′-UTRs), promoter regions, and in noncoding regions (introns).
In some of these diseases, it may be the DNA that forms the G-quadruplex, but in most cases, it is the RNA transcript that forms the G-quadruplex. Here are some of those diseases:
8a. Fragile X Syndrom or FXMR syndrome – a CGG triplet repeat expansion in exon #1 of FMR-1 gene that forms a G-quadruplex
FXMR syndrome is the single most common inherited cause of mental retardation. There is a triple repeat in the first exon of the Fragile X Mental Retardation gene (FMR-1 gene) which does not cause mental retardation in carriers where the repeat is less than 200 nucleotides in length. However, the repeat expands to as many as 2,000 nucleotides in individuals afflicted with FXMR, or Fragile X syndrome. Along with expansion of the “CGC repeat”, there is hypermethylation of the cytosine residues in this area, which results in suppression of the FMR-1 gene. This results in the delay in replication of cells in those with Fragile X syndrome.
Thus the DNA triplet repeat expansion prevents DNA replication in Fragile X syndrome. Interestingly, a functional copy of the helicase enzyme called “Werner syndrome helicase” (aka WRN) can overcome this replication block. (In Werner’s syndrome, the helicase gene is nonfunctional). This led to the discovery of proteins that “unfold” G-quadruplexes, such as WRN. Unlike the effects of the small molecule stabilizers of G-quadruplexes in oncogenes, these small molecules that interact with G-quadruplexes destabilize the G-quadruplexes in Fragile X syndrome. Obviously there is much more to learn about these structures.
References:
1991 Instability of a 550-base pair DNA segment and abnormal methylation in fragile X syndrome
1993 Association of fragile X syndrome with delayed replication of the FMR1 gene
8b. Werner’s syndrome involves a mutation in a G-quadruplex “unfolding protein” called “helicase”
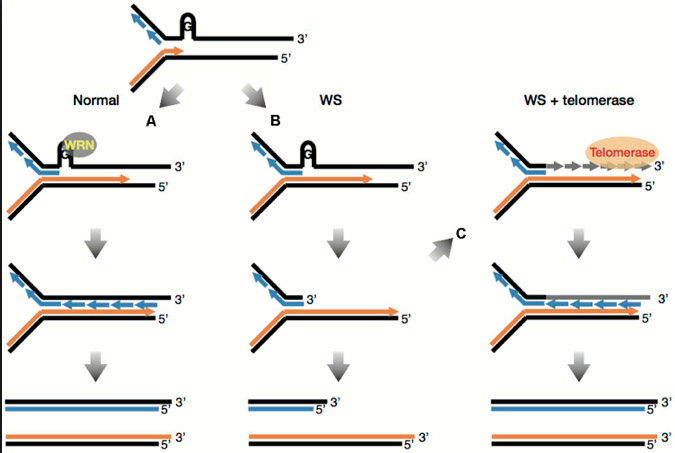
This was a major discovery about the importance of G-quadruplexes. The helicase gene that is mutated in Werner’s syndrome is a critical protein for DNA replication, repair and telomere maintenance. This helicase gene is a member of the RecQ helicase family of enzymes. Mutations in other members of this family also cause disease, such as Bloom syndrome and Rothmund-Thomson syndrome. RecQ helicases are often called the “guardian angels of the genome.” That is quite a strong statement in the 21st century, since it is very hard to see guardian angels today. Although there are many other functions of the Werner’s syndrome helicase, it has an important role in unwinding or unfolding G-quadruplex structures. During replication, G-quadruplexes on the lagging telomere is “unwound” by WRN helicase, making it possible to complete the DNA replication of the lagging strand of DNA. Here is an illustration of the role of WRN helicase in unwinding G-quadruplexes at the telomere.
Image source: 2015 Werner Syndrome-specific induced pluripotent stem cells: recovery of telomere function by reprogramming
8c. Friedrich’s Ataxia – a GAA triple repeat expansion in intron #1 of the Frataxin gene impairs transcription elongation
Unlike the triplet repeat in Fragile X syndrome, the triplet repeat that expands in Friedrich’s ataxia is NOT a G-rich triplet. Instead, it is a “A-rich triplet” of GAA repeats. Nevertheless, the story is similar – the triplet repeat does not cause any disease in carriers with short expansions, whereas when this expansion grows, the individuals display a terrible form of ataxia that was described by Friedrich long ago. The exact method by which this “non-G-rich expansion” causes the disease is still being worked out. What is clear is that the expression of the Frataxin gene is reduced when this expansion gets bigger.
References:
1996 Friedreich’s ataxia: autosomal recessive disease caused by an intronic GAA triplet repeat expansion
2000 Unexpected formation of parallel duplex in GAA and TTC trinucleotide repeats of Friedreich’s ataxia
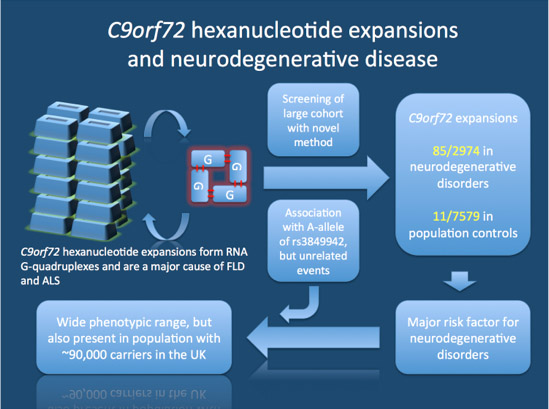
8d. RNA G-quadruplexes form in the C9orf72 inherited form of FTD/ALS
Image source C9orf72 expansions and neurodegenerative disease
“ Study by Beck and colleagues. C9orf72 expansions are thought to produce so-called RNA G-quadruplexes, stable secondary structures that may interfere with normal cellular functions. The authors developed a novel method for rapid screening for these expansions and investigated both a group of patients with various neurodegenerative disorders and population controls. They find these expansions to be present in a more diverse group of patients than previously anticipated. Also, up to 0.2% of the population are carriers of these massive expansions.”
9. DNA methylation and oxidation near G-quadruplex structures may reduce their stability and contribute to aging. Some have proposed that folate and antioxidants may make a major difference in the stability of G-quadruplexes, although no experimental evidence exists to support this claim. References:
2015 G-quadruplexes: A possible epigenetic target for nutrition.
2014 FANCJ promotes DNA synthesis through G-quadruplex structures
“G4 sequences are prone to mutations particularly upon replication stress or in the absence of specific helicases. To investigate how G-quadruplex structures are resolved during DNA replication, we developed a model system using ssDNA templates and Xenopus egg extracts that recapitulates eukaryotic G4 replication. Here, we show that G-quadruplex structures form a barrier for DNA replication. Nascent strand synthesis is blocked at one or two nucleotides from the G4. After transient stalling, G-quadruplexes are efficiently unwound and replicated. In contrast, depletion of the FANCJ/BRIP1 helicase causes persistent replication stalling at G-quadruplex structures, demonstrating a vital role for this helicase in resolving these structures. FANCJ performs this function independently of the classical Fanconi anemia pathway. These data provide evidence that the G4 sequence instability in FANCJ(-/-) cells and Fancj/dog1 deficient C. elegans is caused by replication stalling at G-quadruplexes.”
Reference: 2014 The repair of G-quadruplex-induced DNA damage
“G4 DNA motifs, which can form stable secondary structures called G-quadruplexes, are ubiquitous in eukaryotic genomes, and have been shown to cause genomic instability. Specialized helicases that unwind G-quadruplexes in vitro have been identified, and they have been shown to prevent genetic instability in vivo. In the absence of these helicases, G-quadruplexes can persist and cause replication fork stalling and collapse. Translesion synthesis (TLS) and homologous recombination (HR) have been proposed to play a role in the repair of this damage, but recently it was found in the nematode Caenorhabditis elegans that G4-induced genome alterations are generated by an error-prone repair mechanism that is dependent on the A-family polymerase Theta (Pol θ). Current data point towards a scenario where DNA replication blocked at G-quadruplexes causes DNA double strand breaks (DSBs), and where the choice of repair pathway that can act on these breaks dictates the nature of genomic alterations that are observed in various organisms.”
10. Some (but not all of the Helicases) unwind the 3D structure of DNA in G-quadruplexes. These helicases play a major role in health and aging.
The BRCA1-associated FANCJ Helicase can unwind G4 quadruplexes in vitrond this “G4 resolving function” helps protect DNA and improves genomic stability. When the FANCJ gene is mutated, ” replication stalling” occurs and the genome is vulnerable to oxidation during the stalled replication. As a result, individuals with mutations in the FANCJ gene develop breast cancer or Fanconi’s anemia. Reference: 2013 Specialization among iron-sulfur cluster helicases to resolve G-quadruplex DNA structures that threaten genomic stability
“G-quadruplex (G4) DNA, an alternate structure formed by Hoogsteen hydrogen bonds between guanines in G-rich sequences, threatens genomic stability by perturbing normal DNA transactions including replication, repair, and transcription. A variety of G4 topologies (intra- and intermolecular) can form in vitro, but the molecular architecture and cellular factors influencing G4 landscape in vivo are not clear. Helicases that unwind structured DNA molecules are emerging as an important class of G4-resolving enzymes. The BRCA1-associated FANCJ helicase is among those helicases able to unwind G4 DNA in vitro, and FANCJ mutations are associated with breast cancer and linked to Fanconi anemia. FANCJ belongs to a conserved iron-sulfur (Fe S) cluster family of helicases important for genomic stability including XPD (nucleotide excision repair), DDX11 (sister chromatid cohesion), and RTEL (telomere metabolism), genetically linked to xeroderma pigmentosum/Cockayne syndrome, Warsaw breakage syndrome, and dyskeratosis congenita, respectively. To elucidate the role of FANCJ in genomic stability, its molecular functions in G4 metabolism were examined. FANCJ efficiently unwound in a kinetic and ATPase-dependent manner entropically favored unimolecular G4 DNA, whereas other Fe-S helicases tested did not. The G4-specific ligands Phen-DC3 or Phen-DC6 inhibited FANCJ helicase on unimolecular G4 ∼1000-fold better than bi- or tetramolecular G4 DNA. The G4 ligand telomestatin induced DNA damage in human cells deficient in FANCJ but not DDX11 or XPD. These findings suggest FANCJ is a specialized Fe-S cluster helicase that preserves chromosomal stability by unwinding unimolecular G4 DNA likely to form in transiently unwound single-stranded genomic regions.”
References: 2008 FANCJ helicase defective in Fanconia anemia and breast cancer unwinds G-quadruplex DNA to defend genomic stability
2014 FANCJ promotes DNA synthesis through G-quadruplex structures
Mitochondrial DNA also forms G-quadruplexes in the mitochondria.
These G-quadruplexes correlate well with common sites for deletions that often occur in the mitochondrial DNA. A mitochondrial Helicase called “Twinkle Helicase” unwinds these G-4 structures to allow mitochondrial DNA to be replicated. The Twinkle Helicase is transcribed from a nuclear gene called “C10orf2”. A well known mitochondrial G-quadruplex found in the mitochondrial genome is associated with deletions at this location in renal cell carcinoma. Thus research is now emerging that suggests that mitochondrial genomic Instability maybe due to deletions induced by stalled replication at G-4 structures due to inefficient unwinding by Twinkle. References: 2014 Association of G-quadruplex forming sequences with human mtDNA deletion breakpoints
- G-quadraplexes in TERRA and telomeric DNA may play a role in regulating telomere length
TERRA is a long noncoding RNA that is transcribed from the antisense strand of telomeric DNA. As a consequence, the hexanucleotide repeat of TERRA is “UUAGGG”. The guanine ribonucleotdies of TERRA can form RNA G-quadruplexes, much like how G-rich DNA sequences can form G-4 structures with a cation and Hoogsteen bonds between the guanine ribonucleotides.
TERRA is plays a physiological role and a pathological role in regulating telomerase by interfering with the RNA template used by hTERT to add hexanucleotide repeats to telomeric ends. In addition to this role of TERRA, recent evidence has emerged about the G-quadruplexes in TERRA and the G-quadruplex structures of telomeric DNA. Thus TLS/FUS is both an “RNA and DNA G-quadruplex binding protein”. In the C-terminal region of TLS/FUS, there is an Arg-Gly-Gly domain that binds to both RNA or DNA G-4 structures. The theory is that G-quadruplexes form a scaffold for TLS to bind to RNA and DNA. This then allows TLS to regulate telomere length by histone modifications. If this theory is correct, it points to an active role of a G-quadraplex structure that goes beyond simply blocking DNA transactions.
References:
2013 Regulation of telomere length by G-quadruplex telomere DNA- and TERRA-binding protein TLS/FUS
2014 DNA and RNA quadruplex-binding proteins
13. Both DNA and RNA G-4 structures may regulate estrogen receptor alpha (ER-a) expression.
Since many cancers utilize the ER-alpha receptor as a driver of cancer growth, intense interest has been focused on how the ER-a gene expression is regulated. Estrogen hormone levels, epigenetic regulation, and lncRNA regulation of the ER-alpha gene do not fully account for the variations in ER-a levels in normal cells or neoplastic cells. For this reason, G-Quadruplex mediated gene regulation has been proposed as a possible explanation for the differences in ER-a protein seen in different tissues. 20 G-rich sequences have been found in the ER-a gene, including 3 in the exons of the ER-a gene. G-Quadruplexes have been identified by CD, UV, and NMR spectroscopy. One G-Quadruplex in particular, called the “exon C G-quadruplex”, has been shown to form a very stable DNA G-quadruplex in living cells and decreases ER-a gene expression. This G-rich region is 22 nucleotides long. When a GGG region in this 22nt region was mutated into a AAA, there was a 15-fold increase in the expression of the ER-a gene in bovine cells.
In another experiment, a region in the 5′-UTR of the transcribed ER-a messenger RNA has been found to form a G-quadruplex. Although this G-quadruplex in the 5′-UTR region of the ER-a mRNA does not code for any amino acid, it clearly regulates the rate at which the mRNA transcripts are translated. When cloned and placed in front of a Luciferase reporter gene, this 5′-UTR G-quadruplex reduces expression of the Luciferase by 6-fold!
In summary, both DNA G-4 structures and mRNA G-4 structures exist in the normal expression of the ER-a gene and may regulate transcription (the exon C DNA G-Quadruplex) and translation (the 5′-UTR mRNA G-Quadruplex).
References:
2009 Repression of translation of human estrogen receptor alpha by G-quadruplex formation
2014 DNA and RNA quadruplex-binding proteins
2012 book New Models of the Cell Nucleus: Crowding, Entropic Forces, Phase Separation, and Fractals
2012 book Therapeutic applications of quadruplex nucleic acids
- mRNA G-Quadruplex structures in the 5′-UTR noncoding portion of RNA transcripts
Whereas the early studies of G-quadruplexes was confined to DNA G-4 structures, there has been a recent surge of interest in RNAG-Quadruplexes, since they have been found in vivo. The most common site where G-4 structures exist in RNA is in the 5′-UTR region of the mRNA
Reference: 2012 5′-UTR RNA G-quadruplexes: translation regulation and targeting
“RNA structures in the untranslated regions (UTRs) of mRNAs influence post-transcriptional regulation of gene expression. Much of the knowledge in this area depends on canonical double-stranded RNA elements. There has been considerable recent advancement of our understanding of guanine(G)-rich nucleic acids sequences that form four-stranded structures, called G-quadruplexes. While much of the research has been focused on DNA G-quadruplexes, there has recently been a rapid emergence of interest in RNA G-quadruplexes, particularly in the 5′-UTRs of mRNAs. Collectively, these studies suggest that RNA G-quadruplexes exist in the 5′-UTRs of many genes, including genes of clinical interest, and that such structural elements can influence translation. This review features the progresses in the study of 5′-UTR RNA G-quadruplex-mediated translational control. It covers computational analysis, cell-free, cell-based and chemical biology studies that have sought to elucidate the roles of RNA G-quadruplexes in both cap-dependent and -independent regulation of mRNA translation. We also discuss protein trans-acting factors that have been implicated and the evidence that such RNA motifs have potential as small molecule target. Finally, we close the review with a perspective on the future challenges in the field of 5′-UTR RNA G-quadruplex-mediated translation regulation.”
Here are a few practical phytosubstance-related aspects to the discussion:
15. Quercetin is a strong binder to G-Quadruplex DNA and RNA structures!
This is fascinating and may explain some of the reasons why Quercetin induced apoptosis in the Mayo Clinic study earlier this year.
Reference: 2013 Aminoglycosylation Can Enhance the G-Quadruplex Binding Activity of Epigallocatechin
16. Epigallocatechin binds to G-Quadruplexes. Derivatives of EGC bind even stronger than EGC, such as glucosaminosudes of pentamethylated EGC.
This modified EGC has both an affinity for DNA and RNA G-Quadruplexes.
Reference: 2013 Aminoglycosylation Can Enhance the G-Quadruplex Binding Activity of Epigallocatechin
17. EGCG and Theaflavin both have been found to bind to histone proteins in the cell as well as G-Quadruplexes in the cell
Overall Summary
G-quadruplexes are 3D structures formed by 4 guanines from 1, 2, or 4 strands of DNA or RNA that violate the most fundamental tenant of the “Watson-Crick” dogma of DNA (i.e. that guanine only forms a base pair with cytosine). This “rule violation” has disrupted most of the conventional wisdom about the tertiary structure of DNA. The key structural feature of these G-qaudruplexes is the presence of a monovalent cation (K+ or Na+) in the center of the G-quadruplex. These G-quadruplexes can be “stacked” to form 2, 3, 4, or many stacks. The more stacks there are, the more stable these structures are and the more difficult it is for them to be unfolded. Both the formation and the unfolding of G-quadruplexes are facilitated by specialized G-quadruplex binding proteins that have evolved over millions of years.
Although G-quadruplex structures are mostly found in eukaryotes, they have also been found in some prokaryotes. As many as 376,000 putative G-quadruplex structures could form in the human genome, although this estimate was made purely based on computer modeling. Another 3,000 putative RNA G-quadruplex-forming elements have been identified in the human genome. These RNA G-quadruplex structures are transcribed from 5′-UTR segments of genes.
G-quadruplex structures play a critical role in telomere stably of the 3′-overhang part of the very end of the telomere. They associate with Shelterin proteins and are critical in “G-quadruplex capping of the telomere”. These G-quadruplexes are regulated by BRCA1 gene. The role that BRCA1 gene plays in stabilizing G-quadruplexes may be even more important that its role in double stranded DNA break repair.
The G-quadruplex stabilizing role of BRCA1 on the estrogen receptor gene promoter may explain why gene mutation carriers have such a high incidence of breast and ovarian cancer (i.e. BRCA1 is not turning off the estrogen receptor gene). G-quadruplexes also form in Fragile X Syndrome within exon #1 and in Fredrich’s ataxia in intron #1. However, the triplet repeat expansions in these two inherited genes are the true reason why these patients develop the disease, not the formation of G-quadruplexes per se.
G-quadruplex unfolding is a key role of the Werner’s syndrome helicase protein. When this gene is mutated, the helicase protein cannot unfold the many G-quadruplexes found throughout the human genome. As a consequence, patients with Werner’s syndrome undergo accelerated aging.
Thus the inability to unfold G-quadruplexes may be a major cause of premature aging. There is some evidence that the inability to unfold G-quadruplexes may play a role in normal aging as well.
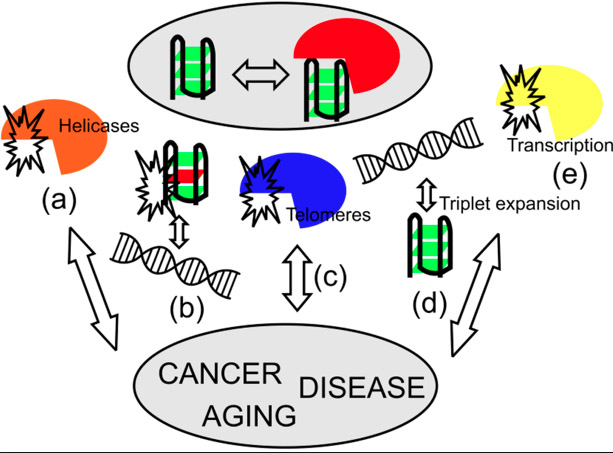
Last of all, G-quadruplexes play a crucial role in carcinogenesis. G-quadruplexes have been found in endonuclease hypersensitivity areas in the promoters or near the promoters of many important oncogenes, including c-Myc, c-kit, bcl-2, KRAS, VEGF, HIF-1a, and other oncogenes. Small molecules that stabilize these G-quadruplexes often induce apoptosis in cancer cells. Here is a diagram that summarizes the many roles of G-quadruplexes in health and disease:
Illustration Reference: Image source DNA and RNA Quadruplex-Binding Proteins
18. Thoughts on G-quadraplexes and evolution
I am fascinated with this wild new 3D structure of DNA. I remember reading about G-quadruplexes when I did a write up on telomeres and TERRA, but really did not understand how fundamental these structures were in Nature. Then when the word came up as I was reading about C9orf72 repeat expansions and familial FTD/ALS, I decided I had to figure out what these structures were.
Now I am convinced that G-quadruplex structures form spontaneously in G-rich regions of DNA and RNA when the weather cools. This could have evolved long ago as the first way of turning off genes at night (cooler temps) or in the winter. Also, since heat will “unwind G-quadruplexes,” they could have evolved as a simple way to initiate transcription of the rising sun (I.e. Circadian rhythms) or turn on growth genes like c-Myc when the weather became warmer in the spring/summer months.
My guess is that simple organisms like prokaryotes had little control over the formation/deformation of G-4 structures that simply turned genes on/off based on ambient temperature, Whereas in more complicated cell (eukaryotes) evolution produced “G-4 folding proteins” and “G-4 unfolding proteins” which could regulate G-4 structures with much tighter transcriptional control. It is also clear to me that the “inability to unwind G-4 structures” is the reason for the accelerated aging phenotype seen with Werners Syndrome. This emerging science may also lead to a better understanding normal aging. The analogy here would be HGPS (Hutchinson-Gilford progeria syndrome). Once upon a time, we thought that HGPS had no similarities to normal aging. Then we found out that progerin forms in skin with normal aging; due to long UV light exposure (UVA) triggering the same alternative splice site in the LMNA gene that is constitutively activated in HGPS! In other words, I bet we will find a G-quadruplex unwinding problem occurs with normal aging, just like it causes accelerated aging in Werner’s Syndrome. Thus WRN mutations causing Helicase dysfunction may provide the clue in normal aging.
Final comments by Vince Giuliano
The question could be asked: “Are-G-quadraplexes 1. pesky artifacts created by Hoogsteen hydrogen bonding that evolution has had to work itself around, or 2. are they tools for the fine-tuning of key biological processes.” I give my usual answer which is “probably both” and also add “but certainly the second.” G-quadraplexes plus the helicase proteins that unwind them are capable of blocking or permitting key DNA transactions in a highly circumstance-driven manner, including gene transcription. As such, they provide another level of cell regulation. beyond that provided by many other mechanisms including histone acetylation, DNA and histone methylation, various species of noncoding RNA, etc. Further, it appears that G-quadraplexes can play other roles besides blocking DNA transactions, such as functioning as a scaffold for the telomere-binding protein, TLS, to regulate telomere length by histone modifications. G-complexes in UTR non-coding portions of RNA point to additional regulatory functions. I think that focusing on regulatory functions of G-quadraplexes is an emerging viewpoint in the literature. Evolution has embraced G-quadraplexes and put them to work. For example, the 2014 publication DNA and RNA Quadruplex-Binding Proteins comments:
“The large number of potential quadruplex structures in all genomes pointed to their importance in cell regulation. Epigenetic modifications and alternative DNA structures appear to provide a higher level of information which may determine and fine-tune complex biological processes at the molecular level. Local DNA structures including cruciforms, triplexes and quadruplexes are often formed in the domains of negatively supercoiled DNA and they could be stabilized and regulated by protein interaction. Since these structures could also be the source of genomic instability, they have to be tightly regulated especially during DNA replication. Telomeric quadruplexes can contribute to the protection of the chromosomal ends. G-quadruplexes in promoter regions can also influence transcription efficiently. Association of quadruplexes with oncogenic and tumor suppressor proteins suggests that quadruplexes may play roles in cancer development and are possible targets for gene therapy. Quadruplex-binding proteins can be divided into several categories. In addition to a well characterized group of proteins which bind specifically to telomeric DNA, we have further classified quadruplex-binding proteins into those which bind to DNA quadruplexes, and those which associate with RNA quadruplexes (see Table 1). Using a new computational tool for examination of conserved G-quadruplex motifs, a great deal of G-quadruplexes conserved across species was identified [145]. Stability of the quadruplexes in evolution suggests the significance of these structures. A deeper understanding of the processes related to their formation, function and recognition will be an important piece of the puzzle in providing better insight into the regulation of living organisms.”


Pingback: AGINGSCIENCES™ – Anti-Aging Firewalls™ has posted a new item, ‘G-qaudruplexes’ – icevckv9