By James Watson (with editorial help from Vince Giuliano)
This is the Part 2 of a three-part series of blog entries on the epigenetic’s of cancer and aging and how those two deadly dragons can be seriously slowed down or stopped with the assistance of plant polyphenols. The Part 1 blog entry tells the central story. It 1. identified similarities in the biological processes and epigenetic’s of cancer and aging, 2. identified therefore how common strategies might be found that address both cancer and aging. 3. described the process of Xenohormesis whereby stress response chemicals developed over millions of years in plants that respond to stresses and keep plants healthy can do the same in humans. 4. provided molecular explanations for the “causality” of cancer and aging, 5. described the processes in cancer and aging of epigenetic silencing of “good” genes and epigenetic activation of “bad” genes, 6. identified a 3 tiered “Pyramid” approach for chemoprevention of aging and cancer, 7. Iidentified the exact interventions involved in each layer of the Pyramid, and 8. Identified how the interventions in the three layers of the Pyramid can be integrated together. This Part 2 blog entry is concerned in more detail with the silencing of good genes in aging and cancer and how polyphenols can prevent that. It lists naturally occurring phytochemicals that inhibit pathways that are known to be unregulated or dysregulated in cancer. The Part 3 blog entry in this Two-Dragon series is concerned: a. the “unsilencing of bad genes” with sirtuin decline and the harmful results in aging and cancer, and B. providing a master list of drugs and natural compounds for cancer chemoprevention. The Part 2 and Part 3 blog provide a series of appendices to the Part 1 entry, They’re published separately because of blog length considerations and because they are of interest in their own right.
Part of Aging and Carcinogenesis is due to gene silencing: histone deacetylation, DNA methylation, & chromatin structure Δ: euchromatin è heterochromatin
We now know that in many cases, gene inactivation is NOT due to a bona fide DNA mutation. Instead, the non-mutated gene can be silenced by an epigenetic mechanism that involves histone deacetylation of the histone proteins surrounding active genes (euchromatin), as well as DNA methylation of a cytosine-rich region of promoter sites (“start location”) on the gene. These cytosine-rich areas are called CpG islands and the cytosine bases can be methylated by the enzyme DNMT. Thousands of “good genes” must remain in the euchromatin state to allow for gene expression to occur. When histones are acetylated, transcription factors can reach the DNA because of the “open structure” of the histones. When the promoter region has a hypomethylated CpG island, the transcription factor can bind to the DNA promoter region. Transcription factors migrate into the nucleus in response to various cellular signaling pathways. are “silenced”, they cannot be turned on by cellular signals, even if the gene is not mutated.
The following discussion explains how gene silencing occurs and then gives some specific examples of gene silencing in aging and cancer. Further, it goes on to illustrate how specific plant polyphenols can be used to unsilence anti-aging and anti-cancer genes and to silence pro-cancer and pro-aging genes.

|
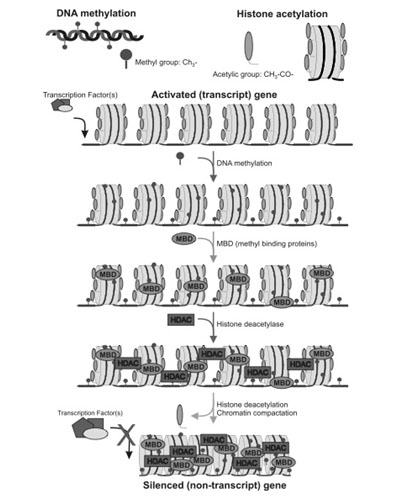
The Sequential Steps in Epigenetic Gene Silencing CpG island hypermethylation Gene silencing by epigenetic mechanisms is an ordered series of events that normally starts with the methylation of cytosine residues. Scientists studying gene silencing have observed two distinct kinds of CpG methylation: one that is age-related that occurs in all tissues as a function of aging, and a cancer-related pattern of methylation that only occurs in cancerous cells. It is important to point out that with aging, there is a global loss of methylation at sites other than CpG islands. The hypermethylation of CpG islands is the exception to this universal decline in DNA methylation with aging. This can be explained in part by the fact that there are different DNA methyltransferases for these different regions of the genome. DNMT1 is responsible for the maintenance of hemimethylated DNA, whereas DNMT3a and 3b are responsible for the methylation of unmethylated DNA. For this reason, it appears that DNMT3a and 3b are responsible for gene silencing. Methyl-binding proteins and Histone deacetylases (HDACs) After the CpG residues have been methylated by DNMT, methyl binding proteins attach to the methyl groups, creating the first layer of protein-based silencers. Methyl binding proteins then recruit histone deacetylases (HDACs) to the histone proteins. Histone deacetylation and chromatin compaction Once the HDACs have been recruited, they remove specific acetyl groups from the lysine side chains on the 9th amino acid of the Histone 3 protein. (H3K9 deacetylation). Once the histone groups have been removed, the nucleosomes can be compacted into a tight configuration called heterochromatin. At this point, the gene is effectively silenced and reactivation of the gene is difficult. |
Image source
Preventing Histone-based Gene Silencing : Inhibiting histone deacetylation of Lysine 9 on Histone 3 (H3K9) with polyphenols
Aging and cancer exhibit epigenetic silencing of genes that should NOT be silenced. Since only 50% of genes have CpG islands at their promoter sites, DNA methylation is not as common a method of gene silencing as histone-based gene silencing, which can occur for almost all genes. Histone-based gene silencing is best understood as an imbalance in the equilibrium between histone acetylation (by HAT) and histone deacetylation (by HDACS). The diagram below illustrates this and how plant polyphenols can inhibit the equilibrium both ways, since some of the polyphenols are both HDAC inhibitors and HAT inhibitors. HDAC inhibition is a way of preventing gene silencing of euchromatin, whereas HAT inhibition is primarily a way of preventing the activation (expression) of repetitive DNA, which normally is silent. The transcriptional activation of repetitive DNA is increasingly being studied since it seems to be the consequence of the global decline in DNA methylation that occurs with aging.

|
How to Prevent Histone Deacetylation The figure above shows the representation of histone modifications (acetylation and deacetylation) by the phytochemicals derived from different food source. Phytochemicals like EGCG, genistein and curcumin play important role in inhibition of histone acetylation by inactivating histone acetyl transferase enzyme. Some other phytochemicals like sulforaphane, curcumin, genistein, phenyl isothiocynate, organosulfur compound, resveratrol and indol-3-carbinol inhibits the deacetylation of relaxed chromatine by inactivating histone deacetylase enzyme. Here is a list of several other phytochemicals that alter histone modifications which are not covered in this figure:
|
Preventing DNA methylation-based Gene Silencing: Inhibiting DNA methylation at the CpG islands found in promoter sites with polyphenols
50% of the promoter sites in human genes have a region that is rich in cytosine bases. This region is called a “CpG island” and is important in the transcriptional activation of gene expression by transcription factors. In genes that have a CpG island (50% of the total), gene silencing can occur by methylation of the cytosine bases in the CpG island. This prevents gene transcription. The enzyme responsible for methylating the cytosines is called DNA methyltransferas. This is normally the first step in gene silencing and occurs before histone deacetylation. Many natural products including green tea, soybeans, garlic, tomatoes, watercress, curry powder, broccoli, brussel sprouts, cabbage, red grapes, and olives prevent this step in the silencing of “good genes”. The diagram below illustrates this.

|
How to Prevent DNA methylation DNA methylation is a biochemical process that is essential for the development of higher organism. Some dietary phytochemicals are reported to inhibit the methylation of deoxycytidine at DNA promoter sites. Hypermethylation of cytidine by DNMTs usually results in transcriptional gene silencing and gene inactivation. Several phytochemicals derived from different food source such as: resveratrol from graps and berries, curcumin from turmeric tea phenols from tea leaves, genistein from soybeans, sulforaphane from broccoli, phenethyl isothiocynate from cauliflower, organosulfur compounds from garlic, quercetin from citrus fruits and lycopene from tomato act as dietary inhibitors of DNA methyl transferases and also alter gene expression via epigenetic mechanisms. Here are some other natural DNMT inhibitors: Compound Source Effect
|
Other methods of Gene silencing – Histone protein trimethylation and Polycomb protein co-repressor silencing of genes
There are other mechanisms of epigenetic silencing not covered in this appendix, but should be mentioned for the sake of completeness. When histone protein tails are deacetylated, the lysine side chains are typically methylated. Single methyl groups or two methyl groups are associated with down regulation of gene expression but not silencing. Trimethylation of the histone protein lysine side chains is typically associated with gene silencing. Polycomb proteins are another form of gene silencing that has been evolutionarily conserved from lower organisms that do not methylate DNA. The significance of Polycomb protein-based gene silencing is not as clear has the other epigenetic mechanisms described in this Two-Dragon series of entries. Nevetheless, this Polycomb silencing is important to understand and occasionally comes up in discussions of gene silencing.
Gene Silencing by non-coding RNA – miRNA silencing of tumor suppressor genes and activation of oncogene expression
A third method by which genes can be silences is by the inhibition of messenger RNA (mRNA) by non-coding RNA called miRNA (also called microRNA). These miRNA bind to mRNA, making them incapable of creating a protein, or increasing the degradation of the mRNA, thereby eliminating the possibility that the mRNA can create a protein. This means that the gene may be transcribed (i.e. a messenger RNA is made), but the mRNA transcript is not “read” and translated into a protein in the ribosomes. This method of gene silencing can also be used to activate or increase the expression of an oncogene. Most oncogenes are actually good and necessary genes expressed with normal cellular function, but become overexpressed with cancer. When this occurs, such a gene is referred to as an “oncogene”. The diagram below illustrates how miRNA can inhibit a tumor suppressor mRNA and activate an oncogene. Several naturally occuring foods have active ingredients that prevent tumor suppressor mRNAs from being silenced by miRNA, as well as preventing oncogene mRNAs from being translated into proteins. These natural foods include berries, green tea, soybeans, broccoli, and curry spice.

|
How to Suppress Oncogenes and Activate Tumor Suppressor Genes with Tea, Tofu, Food, and Dietrary polyphenols can affect microRNA (miRNA) expression. miRNAs are noncoding RNAs that regulate gene expression after the gene is transcribed into a messenger RNA (mRNA). miRNA are transcribed in the nucleus into primary miRNA (pri-miRNA) which is further cleaved by Drosha into precursor miRNA (pre-miRNA). Pre-miRNA is exported from nucleus to the cytoplasm and further processed by Dicer into miRNA duplex. Single strand of miRNA duplex (also called mature miRNA) leads this complex to mRNA cleavage or translation repression, which is dependent on miRNA:mRNA complementarity. Dependent on various factors, miRNA can have either an oncogenic role (called onco-miRNAs) if the target mRNA is a tumor suppressor gene, or a tumor suppressive role (tumor-suppressor miRNAs) if the target molecule is an oncogene. Dietary polyphenols can impact expression level of miRNAs and participate in gene expression regulation. |
How to “Reactivate” your Silenced Genes
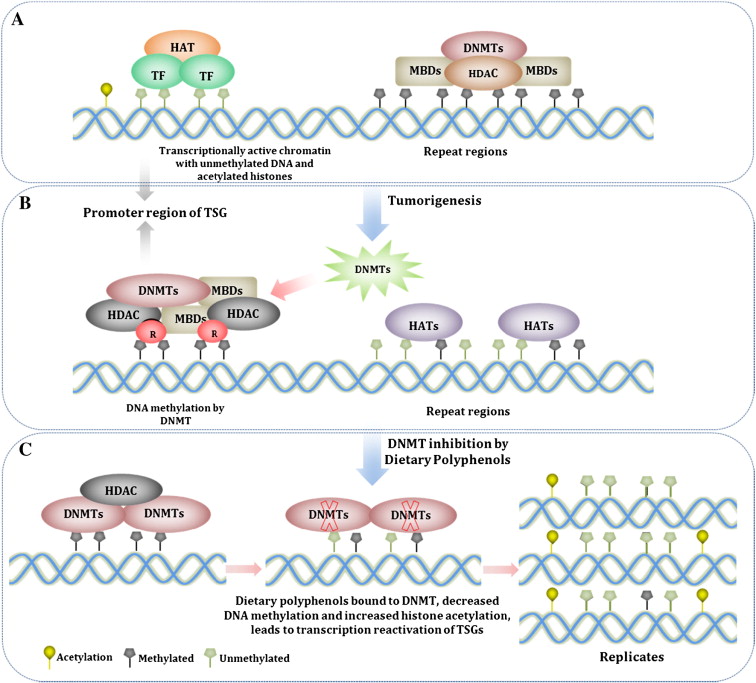
The epigenetic silencing of “good genes” is not irreversible. Natural compounds, when used on a regular daily basis can reverse this phenomena and reactivate silenced tumor suppressor genes. The diagram below illustrates this:
|
How To Turn Your Genes Back On A. Inhibit HDACs – Histone deacetylases are enzymes found in the cell nucleus that remove acetyl groups from the histone proteins. This inactivates the transcriptional machinery. This is the most common way that dietary polyphenols turn genes back on. This step usually precedes DNMT activity. B. Inhibit HAT – Histone acetyltransferases are enzymes that reacetylate histone proteins, opening the chromatin structure of DNA, allowing the genes to be reached by transcription factors. At first glance, you would think that HAT activation would be good, rather than inhibition. Unfortunately, HATs also have a “dark side” to their function, especially with regards to repetitive DNA sequences. Normally, repetitive DNA is silenced and should NOT be transcribed, since this results in genomic instability. Unfortunatley with aging and with cancer, there is a global decline in DNA methylation (opposite of promoter regions where there is an increase in DNA methylation). When HATs bind to demethylated regions of repetitive DNA, they acetylate the repetitive DNA, allowing transcription of the repeat regions. This can contribute to tumorigenesis and genomic instability. There are several natural compounds have been shown inhibit HATs and thereby prevent the “unsilencing” of repetitive DNA. This includes allspice (eugenol, ericifolin), cashew nut oil (anacardic acid), green tea (EGCG) soybeans (genistein), and turmeric (curcumin). C. Inhibit DNMTs – DNMT activity is blocked by dietary polyphenols by forming hydrogen bonds with amino acids (Pro, Glu, Cys, Ser, and Arg) in the catalytic pocket of DNMT. Newly synthesized DNA strands are semi-methylated after the first round of DNA replication and become progressively more demethylated after several rounds of replication due to the dilution effect. This is why life long dietary intake of polyphenols that inhibit DNMT is so important. |
Specific Examples of Genes known to undergo epigenetic silencing
It is important to point out that not all genes have CpG islands. In humans, approximately half of the all genes have CpG islands, but the very important genes involving “housekeeping genes” and “tissue-specific genes” seem to have a disproportionate share of CpG islands. These important genes are silenced by a combination of histone deacetylation and CpG island DNA methylation at the promoter sites. This results in the inability of a transcription factor to access the gene and to bind to the promoter site. With aging, many genes are silenced this way without any mutation of the DNA. There are some specific genes that are silenced via this epigenetic mechanism that play a huge role in aging. Here are some of them:
1. DNA repair gene inactivation by epigenetic silencing – Four examples of this are the genes hMLH1, MGMT, WRN, and BRCA1
hMLH1 – this an important gene for DNA mismatch repair, which is the way that microsatellite-unstable regions are repaired to avoid them becoming bigger. Gene inactivation by CpG island hypermethylation has been observed at the promoter site for this gene
MGMT – this is an important gene for fixing mutant guanine bases that become chemically modified by a methyl or alkyl group. This modification results in the “G” being read as a “A” (i.e. creating a G-to-A single nucleotide polymorphism). MGMT removes the pro-mutagenic guanine residue. Unfortunately, this gene is inactivated by CpG island hypermetylation with aging.
WRN – this is the gene responsible for Werner’s syndrome. This gene codes for the WRN protein which has helicase and exonuclease activity. With aging, the CpG island at the promoter site for this gene is hypermethylated, effectively silencing this gene. As a consequence, the manifestations are a progeroid phenotype and increased risk of cancer due to extreme sensitivity to DNA damaging drugs or toxins.
BRCA1 – this gene is the most common gene that is mutated with hereditary breast cancer. However in sporadic cases of breast and ovarian cancer, the BRCA1 gene can be silenced by CpG island hypermethylation in the promoter site of this gene.
2. Progeroid syndromes and atypical Werner’s syndrome – inactivation by epigenetic silencing. Two examples of accelerated aging genes – Lamin A/C and WRN
Laminin A/C – Mutations in the Laminin A/C gene produce a syndrome called Hutchinson’s Gilford progeria. Although most progeroid syndromes are due to DNA mutations, we now know that a progeroid-like phenomena can be acquired due to epigenetic silencing of the same gene that is mutated in the hereditary form of HG progeria. The Laminin A/C gene codes for two different laminin proteins that make up the scaffolding just inside the nuclear double membrane. When this gene is hypermethylated at the promoter site, a syndrome called “atypical Werner’s syndrome” occurs.
WRN – this is called the “Werner gene” mentioned above. It is mutated in classic Werner’s syndrome (WS). Patients with WS develop accelerated aging and manifest cataracts, type II diabetes, osteoporosis, arteriosclerosis, and cancer at an earlier age and at increased incidence. Cases of WS where the only problem is CpG island hypermethylation at the promoter site site. Again, this is where we need a “DNA methylome” for proper diagnosis.
3. Alzheimer’s disease – Epigenetic “unsilencing” of PS1 and BACE genes. Alzheimer’s disease appears to be a multifactorial problem that is age-related. Epigenetic silencing is only one of the explanations for the disease. Several explanations that involve epigenetic mechanisms may be involvedwith AD – homocystein/SAM dysregulation, DNA methylation/demethylation of genes, and the effects of folate and B12 deficiencies on the methylation status of genes involving AD.
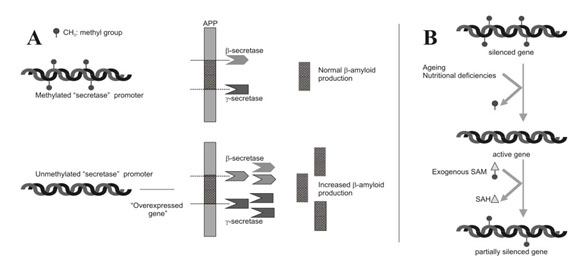
Image source
|
Two Epigenetic Explanations for Alzheimer’s Several studies have shown that both the gamma-secretase gene and the beta-secretase genes are under the control of DNA methylation of their promoter sites. Beta-secretase cleaves APP in the wrong place, creating amyloid-beta. DNA demethylation causes the overexpression of PS1 and BACE genes, which result in an increase in amyloid-beta. Nutritional deficiencies can also explain AD. In vitro studies of neuroblastoma cells show that nerve cells cultured with a B12 or folate deficiency result in the over production of amyloid-beta. Adding B12 or folate can reverse the increase in amyloid-beta. Exogenous SAM can also restore the methylation pattern and silence the gene in a physiologic way. |
Naturally occurring phytochemicals that inhibit pathways that are known to be unregulated or dysregulated in cancer.
Big Pharma is developing drugs to block most all of these pathways. You can block them yourself by going for a walk in the jungle, the forests, or the mountains.
You may notice a recurrent theme in the following table: the best health-producing natural plant substances may be found in the dumpsters of industrial plants that produce commodity food and drink substances like orange juice, wines, tomato ketchup, olive oil and applesauce. This is because the most-powerful naturally-produced stress resistant phytochemicals substances occur in the skins and rinds that are normally removed in the course of making commodity products. Skins are where oranges, grapes, tomatoes, olives, grapefruit, nuts, apples, and many other plants foods have found an evolutionary need to concentrate their chemical stress defenses. If we want to take advantage of the xenohormetic properties of these substances we have to find them where they are – today often in the dumpster.
Pathway Inhibited (or activated) — Compound Natural Sources of Compound
Nonspecific Tyrosine Kinase Inhibitors
genistein — soybeans – maybe we can get these from soy milk manufacturing plants
resveratrol — polygonum cuspidatum roots, Japanese knotweed, red grape skins
quercetin — apple skins, onions, etc. – maybe we can get these from apple juice plant garbage
luteolin — tomato skins – maybe we can get these from Campbell’s tomato soup plant
erbstatin — Streptomyces extract
itaconic acid — Aspergillus niger, Aspergillus terreus extract
staurosporine — Streptomyces staurosporeus
beta-sitosterol — saw palmetto, avocado, pumpkin seeds, cashew fruit, dandelion coffee, etc.
daucosterine a beta-sitosterol — isolated from roots of Acanthopanax sessiliflorum
macrostemososide A — ginseng
laxogenin — Smilax sieboldi
chinenoside II — Allium chinense (rakkyo)
ginsenoside ed — Red Ginseng
icariside — Epimedium grandiflorum, Epimedium sagittatum, E. thunbergianum, etc.
icaritin — Epimedium brevicornu
EGFR tyrosine kinase inhibitors (above list) and:
lavendustin A — streptomyces griseolavendus
herbimycin A — streptomyces hygroscopium
other natural isoflavones
other natural quinolones
Angiogenesis inhibition
procyanidin oligomers
1,25-hydroxyvitamin D ?
emodin — Oxygonum sinuatum
coleon A lactone — Plectranthus barbatus, Triphala churna (3 fruits)
Vitamin E, EPA, DHA — krill oil, fish oil, or myriad algae
mushrooms — “2nds” at mushroom farm
curcumin — “left overs” from curry spice factory
allyl mercaptan — garlic
chloroplast glycoglycerolipids — spinach
proteoglycan molecules — bindweed (Convolvulus arvensis) extract
Aromatase inhibitors
apigenin —
naringenin — citris rinds from orange juice factory garbage
hesperitin — citris rinds from grapefruit juice plant dumpster
mushroom polysaccharides — left-overs or surplus from mushroom farms
5alpha-reductase inhibitors
ECGC — tea plantation extra leaves, white tea surplus, Green tea surplus store
Mushroom extracts — Ganoderma lucidum
Angiogenesis-related tyrosine kinase
hydroxycinnamic acid — cinnamon farms, cinnamon factories
other forms of cinnamic acid
liposterolic extract of Serenoa repens
β-sitosterol
Beta-catenin inhibitors
Wild cherry bark extract — Prunus Serotina
Kras/Braf inhibitors
EPA, DHA — fish oil, krill oil, myriad algae
c-myc inhibitors
prenylated xanthones — mangosteen rinds from the mangosteen fruit packaging plant
garcinol — mangosteen rinds from the mangosteen fruit dumpster behind the juice factory
Selective apoptosis inducers
boswellic acid — Frankincense that the Wise men forgot to take to Bethlehem
in cancer cells: pterostilbene — Red Sandalwood oil on sale at 99 cents store (it’s not selling very well elsewhere), a-santol Red Sandalwood oil
ursolic acid — apple skins from the apple pie factory and the apple juice factory dumpster
diosgenin — Wild Yam farm
charantin (b-stiosteryl) — Bitter Melon farm – no one is buying these melons for dinner
mormodin — Bitter Melon farm – we could get two for the price of one
allyl mercaptan — Garlic from Gilroy, CA – they make so much of it that the whole town stinks!
Vitex — Vitex Agnus-Castus farm
Icarisides — Epimedium grandiflorum, Epimedium sagittatum, E. thunbergianum, etc.
Cellular senescence inhibitors
kinetin — coconut water
trans-zeatin — coconut water
gibberellins — I have no idea where to find these or if they are good for you
polyamines — I have no idea where to find these or if they are good for you
NF-kB inhibitors
curcumin — curry powder factory left-overs
silymarin – milk thistle
Many other phytosubstances — In Vince’s Treatise, he lists 39 substances he takes as supplements that inhibit expression of NF-kappaB
COX2 inhibitors
EGCG, EGC — green tea factory seconds. — white tea store sale
FGF inhibitors — Gentisic acid rinds or skins of lots of fruits
HDAC inhibitors
ericifolin — Allspice factory left overs
anacardic acid — Cashew nut shell oil – you can buy this really cheap
cinnamic acid — cinnamon farm
hydoxycinnamic acid — cinnamon yard sales
procyanidins — cinnamon factory
zerumbone — ginger factory – no one is buying this except in Japan
DNMT inhibitors
chlorogenic acid — green coffee beans
caffeic acid — green coffee beans
EGCG, EGC — green tea
Genistein — soybeans
Heat Shock Protein inhibitors
geldanamycin — Streptomyces hygroscopicus
PICK/Akt?mTOR inhibitors
caffeine (PI3K/Akt/mTOR) — green tea, coffee
xanthines — cocoa (chocolate)
resveratrol (mTOR inhibition) — grape skins, wine
genistein (Pi3K/Akt) — soybeans
quercetin (mTOR) — apple peelings, onions, especially the outer layers
butein (PI3K/Akt) — Toxidodendron vernicifluum (aka Rhus vermocoflua)
capsaicin (PI3K/Akt) — hot chili peppers
deguelin (Akt/mTOR) —roots of Lonchocarpus nicou
tocotrienol (Akt) — Vitamin E
ERK activators
soyasaponins — soybeans
PPaR activators
lycopene, lutein, luteolin — Campbell’s tomato soup plant garbage dumpster
AMPK activators
resveratrol — winery garbage, Welch’s grape juice plant waste, grape skins, Japanese knotweed
nduratin A — Bitter melon extract
Galegene — guanidine derivative
Autophagy activators
apigenins
naringenin — grapefruit rinds
resveratrol — Japanese knotweed, grape skins
caffeine — coffee, tea, etc.
capsaicin — hot chili peppers
FOXO3a activators
n-Tyrosol — white winery
hydroxytyrosol — white winery
secoiridoids — olive oil pressing plant
Nrf2 activators
isothiocyanates — broccoli, brussel sprouts, cabbage, cauliflower
lycopene, lutein — tomato skins
capsaicin — hot chili peppers
pterostilbene — blueberries, grapes
EGCG — green tea
(Nrf2 activators and NF-kappaB inhibitors tend to be the same)
Sirtuin activation
resveratrol — grape skins
butein — Rhus verniclfua
piceatannol food additive
fisetin — strawberries
quercetin — capers, lovage, apples, tea, onions, citrus fruits, green vegetables
Klotho activation
Chinese herbs – selected Chinese herbs
Immune activators
beta 1,3 glucan — fungi cell walls
bacterial DNA — Lactobacillus fermentum extract
—————————————————————————
See the important note on bioavailability of plant polyphenols and nano-particle delivery at the end of the Part 3 blog entry.
DATA DISCLAIMER:
The tables and compilations of data in this and the other blog entries in the Two Dragons series are intended to be illustrative of the main points to be made. They are compiled from various sources, in most cases are incomplete, and may contain occasional errors.
MEDICAL DISCLAIMER: FROM TIME TO TIME, THIS BLOG DISCUSSES DISEASE PROCESSES. THE INTENTION OF THOSE DISCUSSIONS IS TO CONVEY CURRENT RESEARCH FINDINGS AND OPINIONS, NOT TO GIVE MEDICAL ADVICE. THE INFORMATION IN POSTS IN THIS BLOG IS NOT A SUBSTITUTE FOR A LICENSED PHYSICIAN’S MEDICAL ADVICE. IF ANY ADVICE, OPINIONS, OR INSTRUCTIONS HEREIN CONFLICT WITH THAT OF A TREATING LICENSED PHYSICIAN, DEFER TO THE OPINION OF THE PHYSICIAN. THIS INFORMATION IS INTENDED FOR PEOPLE IN GOOD HEALTH. IT IS THE READER’S RESPONSIBILITY TO KNOW HIS OR HER MEDICAL HISTORY AND ENSURE THAT ACTIONS OR SUPPLEMENTS HE OR SHE TAKES DO NOT CREATE AN ADVERSE REACTION
—————————————————————————-





Pingback: PART 1: Slaying Two Dragons with One Stone – How to Prevent Cancer and Aging with the Same Strategy | AGING SCIENCES – Anti-Aging Firewalls
Hi Vincent,
from this two parts article looks like the effective way to inhibit DNMT, HAT, HDAC would be to take curcumin, resveratrol, genistein and EGCC. If in addition to that Lycopene and quercitin is included most of the pathways would be tackled.
Best. Santiago
Intermittent Fasting combined with a ketogenic diet seems to offer the ability to slay the dragons of age, cancer, dementia, high blood sugar and so many ills that it seems to good to be true. However, autophagy seems to help and ketones seem to help and I do not see why we as animals can not benifit from a combination of Intermittent Fasting and a ketogenic diet. Eric
Althou I do continue to keep an eye on new areas like Carbon 60 and Manoheptulose.
Moringa oleofera, part of the human/animal diet in tropical regions, looks like another interesting source of zeatin (among other other things):
http://www.ncbi.nlm.nih.gov/pubmed/17089328 – Moringa oleifera: a food plant with multiple medicinal uses.
Pingback: PART 3: Slaying Two Dragons with the Sound of Silence: – How to Keep Your Repetitive DNA Turned Off with “3 Songs”: Sirtuins, Polycomb Proteins, and DNMT3. And a Master List of Drugs and Natural Compounds for Cancer Chemoprevention | AGING SCIEN
Pingback: Quorum sensing Part 1: quorum sensing inhibition via phytochemicals – a new approach against infectious diseases. | AGING SCIENCES – Anti-Aging Firewalls
Pingback: DEEPER INTO THE EPIGENETICS OF YOUNGING 1.1 - INTERACTIONS OF H3K9, H3K4, H3K27, H3K79 AND BIVALENT HISTONE POST-TRANSLATIONAL MODIFICATION (PMT) DOMAINS - AGINGSCIENCES™ - Anti-Aging Firewalls™AGINGSCIENCES™ – Anti-Aging Firewalls™
Pingback: Healthy, active and productive till 100. Laying out the Adult Aging Process, a Breakthrough and my Personal Story - AGINGSCIENCES™ - Anti-Aging Firewalls™AGINGSCIENCES™ – Anti-Aging Firewalls™