By James Watson (with some help from Vince Giuliano)
This is the Part 3 of a three-part series of blog entries on the epigenetic’s of cancer and aging and how those two deadly dragons can be seriously slowed down or stopped with the assistance of plant polyphenols. The Part 1 blog entry tells the central story. It 1. identified similarities in the biological processes and epigenetic’s of cancer and aging, 2. identified therefore how common strategies might be found that address both cancer and aging. 3. described the process of Xenohormesis whereby stress response chemicals developed over millions of years in plants that respond to stresses and keep plants healthy can do the same in humans. 4. provided molecular explanations for the “causality” of cancer and aging, 5. described the processes in cancer and aging of epigenetic silencing of “good” genes and epigenetic activation of “bad” genes, 6. identified a 3 tiered “Pyramid” approach for chemoprevention of aging and cancer, 7. Iidentified the exact interventions involved in each layer of the Pyramid, and 8. Identified how the interventions in the three layers of the Pyramid can be integrated together. The Part 2 blog entry is concerned in more detail with the silencing of good genes in aging and cancer and how polyphenols can prevent that. It lists naturally occurring phytochemicals that inhibit pathways that are known to be unregulated or dysregulated in cancer. This Part 3 blog entry is concerned with: A) the silencing of bad genes and repetitive DNA to avert harmful consequences in aging and cancer, and B) providing a master list of drugs and natural compounds for cancer chemoprevention. The Part 2 and Part 3 blog entries provide a series of appendices to the Part 1 entry, They are published separately because of blog length considerations and because they are of interest in their own right.
PART A – Silencing repetitive DNA sequences
The Sound of Silence – How Leonard Guarante discovered Sir2, the repetitive rDNA silencer, in Baker’s yeast
A good way to present this is by taking a little romp through the history of actions of sirtuins.
Working with Baker’s yeast at MIT, back from 1997 to in 2004 Dr. Leonard Guarante screened yeast colonies for long-lived strains and discovered that a single mutation in the gene SIR4 caused the protein Sir2 to gather at the most repetitive DNA sequence in yeast – the rDNA repetitive gene sequence where there are about 100 copies of this gene for ribosomal proteins(ref)(ref). Unfortunately, this repetitive area is unstable and the gene copies tend to recombine with each other. This is called “genomic instability” and is seen with many other genes in humans (humans do not have an unstable rDNA area, however). Recombination of repetitive DNA is a major cause of human cancer and other human diseases such as Huntington’s disease(ref)(ref)(ref).
ERCs and gene silencing by Sir2 – How an extra copy of SIR2 gene extended lifespan by 30% & 50%
In yeast, a very unusual event occurs when the rDNA copies combine with each other. With yeast aging, the mother yeast cell starts spinning off circular rings of this rDNA, which “pop out of the DNA”. These DNA rings are called “extrachromosomal rDNA circles” or ERCs(ref). When the mother yeast cell divides, these ERCs are also copied but all the ERC copies stay in the mother yeast cell. With successive cell divisions, the mother yeast cell becomes over-run with all these ERCs. These ERCs cause aging. The cellular energy and resources required to replicate all these ERCs eventually spells death for the mother yeast after about 20 cell divisions. When an extra copy of the SIR2 gene was added to the yeast cells, this extended the lifespan of the yeast by 30%. When the same experiment was done in nematodes, the extra copy of SIR2 extended lifespan by 50%. This was the first indication that SIR2 was a “longevity gene” and that the mechanism of life span extension was “gene silencing” of repetitive DNA(ref). So, in yeast at least something can be done about repetitive DNA and its life-shortening consequences, and that something involves SIR2. Of course, the question arose “What about mammals, including us?”
Histone deacetylation = Gene silencing
Once the Sir2 protein was discovered to be a gene silencer, the molecular mechanisms for this effect on gene expression was discovered to be the removal of acetyl groups from the histone proteins that shroud DNA. It was discovered that this Sir2 histone deacetylator could only do this if NAD+ was present. NAD+ was a cofactor required for decetylation to occur and high levels of NAD+ and low levels of NADH would stimulate Sir2. Since high levels of NAD+ reflected low energy levels in the cell, it was determined that these histone deacetylators were “energy sensors” responding to food (calorie) availability. This was the impetus to do caloric restriction studies in the Baker’s yeast.
Calorie Restriction -> Sir2 Activation -> Longevity
Caloric restriction(ref),(ref ),(ref) had already been shown to extend lifespan in experiments done 70 years prior, but there were still major misconceptions about how caloric restriction could extend lifespan. The “conventional wisdom” at the time of Dr. Guarente’s yeast studies (2004) was that caloric restriction was due to slowing down metabolism, which would then reduce free radical production by mitochondria. (Lane, Ingram, Roth, Scientific American, 2004). Unfortunately, experimental studies did not support this “rate of metabolism theory”, since caloric restriction does not slow down metabolism in yeast, worms, and humans. This lack of experimental evidence for the “rate of metabolism theory” lead Guarante to consider Sir2 as a possible mediator of caloric restriction’s effects on longevity. With yeast caloric restriction activates Sir2 by two different mechanism:
1. PNC1 – PNC1 is an gene that codes for an enzyme that gets rid of nicotinamide, a molecule similar to vitamin B3. Nicotinamide inhibits Sir2, so getting rid of it activates Sir2
2. Respiration – mitochondrial energy production is called “respiration” and has a by-product called NAD+. Lowering NAD+ levels activates Sir2.
The final “proof” that Sir2 is required for the longevity effects of caloric restriction was done by “knocking out” the SIR2 gene, which eliminated the life span extending effects of caloric restriction in yeast. Adding an extra copy of SIR2 gene to fruit flies further extended the life span extending effects of caloric restriction(ref).
Resveratrol -> Sir2 Activation -> Longevity – the claims, the questions, and the vindication
When David Sinclair joined Dr. Guarente’s laboratory at MIT as a post doc, he did the now famous experiment that showed how resveratrol, a polyphenol found in grape skins and wine, activated Sir2 in yeast, which led to world wide attention to Sir2 as the ultimate “longevity gene” and red wine as the “fountain of youth”, explaining the French paradox (The paradox of why French people do not have as much heart disease as would be predicted, based on their lifestyle). Unfortunately, the results of this experiment could not be reproduced in other yeast laboratories and the scientific validity of this conclusion was questioned. Although few were questioning the longevity effect of resveratrol, some were questioning if this effect was mediated by Sir2. Over the past 7 years, this controversy has continued to smolder, with many claiming that the primary mechanism of action for resveratrol is AMPK activation. Earlier this year, however, David Sinclair and Leonard Guarente published a follow-up study that eliminated the questionable reagent used in the experiment. Without the questionable reagent, the results of the resveratrol experiment was the same as the original study. This paper has largely silenced the critics of the silencing ability of resveratrol on repetitive rDNA circle formation(ref).
There is much more to the story of sirtuins than can be conveyed here. You can review the blog entries such as ones cited above and Genomic stability, DNA repair and the sirtuin SIRT6, SIRT3 research – tying together knowledge of aging, What does resveratrol do?, SIRT1, the hypoxic response, autophagy and hormesis, SIRT1, mTOR, NF-kappaB and resveratrol, and Visit with Leonard Guarante. Although some of these are a few years old, they do not predate much of the research just described.
Other Ways To Stimulate your Sirtuins
There are activities, naturally occurring molecules and synthetic molecules that activate Sirtuins. These “sirtuin activating compounds” are called STACs. Here is a list of natural STACS and synthetic STACS. Please note that some activities or compounds only activate the enzyme, whereas other activities/compounds also activate gene expression of the SIRT enzyme:
| Compound or Activity | Source/Mechanism of Action |
| Exercise (E) | E -> SIRT activity, E -> SIRT gene expression, but net effect is overall |
| Caloric Restriction (CR) | CR -> SIRT activity AND ->SIRT gene expression |
| Dietary polyphenols | polyphenols -> SIRT activity but do not -> SIRT gene expression |
| Resveratrol | 4.6-5.2 fold -> in SIRT activity – not affected by lack of vitamin C |
| Fisetin | |
| Quercetin | 2.1-2.5 fold -> in SIRT activity – not affected by lack of vitamin C |
| ECg | 1.85–1.91 fold -> in SIRT activity – not affected by lack of vitamin C |
| EC | 0.99-1.09 fold -> in SIRT activity – not affected by lack of vitamin C |
| EGCg, EGC, myricetin, gallic acid | very weak SIRT activators that loose their effect without vitamin C |
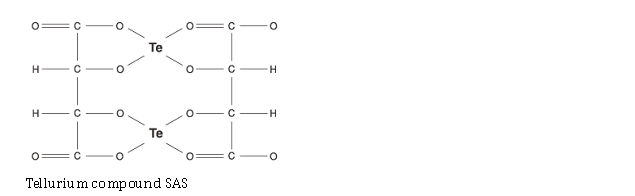
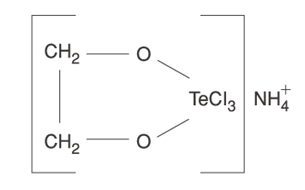
| Tellurium compounds | -> SIRT activity AND -> SIRT gene expression |
| Synthetic analogs of resveratrol | synthetic STACS -> SIRT activity but do not -> SIRT gene expression |
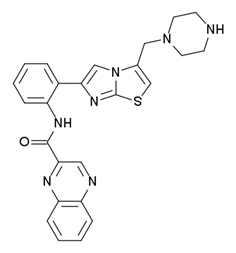
| SRT 1720 | 100X more potent than resveratrol |
| SRT 2183 | in development by Glaxo-Smith-Kline |
| SRT 1460 | in development by Glaxo-Smith-Kline |
| Icarin | -> SIRT via MAPK kinase, does not -> SIRT gene expression |
| 2-Deoxyglucose (2DG) | -> SIRT activity AND ->SIRT gene expression via -> association with redox s |
Some references:
Gurd BJ, Perry CG, Heigenhauser GJ, Spriet LL, Bonen A., High-intensity interval training increases SIRT1 activity in human skeletal muscle, Appl Physiol Nutr Metab. 2010 Jun;35(3):350-7.
Marie Lagouge, Carmen Argmann, Zachary Gerhart-Hines,Hamid Meziane, Carles Lerin, Frederic Daussin, Nadia Messadeq, Jill Milne, Philip Lambert, Peter Elliott, Bernard Geny, Markku Laakso, Pere Puigserver, and Johan Auwerx, Resveratrol Improves Mitochondrial Function and Protects against Metabolic Disease by Activating SIRT1 and PGC-1a, Cell, Vol 127, pp1-14, December 15, 2006.
Meital Halperin-Sheinfeld, Asaf Gertler, Eitan Okun, Benjamin Sredni*, and Haim Y. Cohen, The Tellurium compound, AS101, increases SIRT1 level and activity and prevents type 2 diabetes, Aging, Vol 4(6), pp 436-447, June, 2012
Marie Lagouge, Carmen Argmann, Zachary Gerhart-Hines,Hamid Meziane, Carles Lerin, Frederic Daussin, Nadia Messadeq, Jill Milne, Philip Lambert, Peter Elliott, Bernard Geny, Markku Laakso, Pere Puigserver, and Johan Auwerx, Resveratrol Improves Mitochondrial Function and Protects against Metabolic Disease by Activating SIRT1 and PGC-1a, Cell, Vol 127, pp1-14, December 15, 2006.
Lin Wang, Ling Zhang, Zhi-Bin Chen, Jia-Yong Wu, Xin Zhang, Yun Xu, Icariin enhances neuronal survival after oxygen and glucose deprivation by increasing SIRT1, European Journal of Pharmacology, Volume 609, Issues 1–3, 1 May 2009, Pages 40–44
Qinghong Zhang*†, Su-Yan Wang*, Capucine Fleuriel‡, Dominique Leprince‡, Jonathan V. Rocheleau§, David W. Piston§, and Richard H. Goodman, Metabolic regulation of SIRT1 transcription via a HIC1:CtBP corepressor complex, PNAS, Vol 104(3), pp 829-833, January 16, 2007
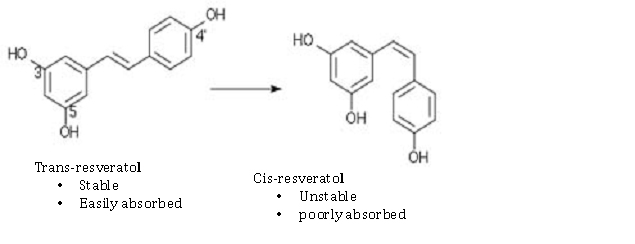
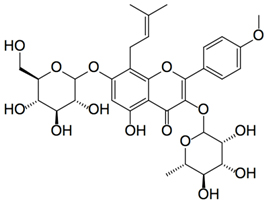
Here are some examples of the chemical configurations of these gene silencing compounds that work on Sirtuin activation or SIRT gene expression. As you can see, there are two enantiomers of resveratrol: trans-resveratrol and cis-resveratrol. The trans form is stable and easily absorbed. The cis form is unstable and poorly absorbed. For this reason, most dietary supplements are made with the trans form of the compound.
Icariin
Differences in Caloric Restriction Mechanisms in Different Species
A very important distinction should be made in humans, where there are 7 different sirtuins. In humans, the sirtuin that is involved with metabolic control is SIRT1, which is the ortholog to the yeast SIR2. Because all of the 7 Sirtuins are “redox sensitive”, however, there will be other effects of caloric restriction besides the control of metabolic pathways. In other words, caloric restriction (CR) has traditionally been studied just with SIRT1 and the effects of SIRT1 on transcription factors. The other Sirtuins (SIRT2, SIRT3, SIRT4, SIRT5, SIRT6, and SIRT) are also “redox sensitive” but have completely different functions/effects. To date, SIRT1 has not been shown to have a longevity effect in humans. However recent studies have found that SIRT6 does have a longevity effect in humans. Another reason for the controversy about Sirtuins is that caloric restriction appears to have different pathways that mediate their effects in different organisms. As you can see below, mitochondria play a crucial role in CR. The following diagram illustrates some of these differences:
Reference: Leonard Guarante, Mitochondria- A Nexus for Aging, Calorie Restriction, and Sirtuins?, Cell, Vol 132(2), pp171-176, January 25, 2008.
Calorie restriction increases life span in all model organisms, but has not shown a longevity effect in humans yet. Two longitudinal primate studies both showed positive effects on health, but only one of the two showed an increased life span effect. The diagram above from Guarante’s review article shows how it the yeast SIR2 gene othologs in mice and humans is the SIRT1 gene, but in nematodes sirtuins do not appear to be involved with the caloric restriction pathway. This may be due in part to the fact that repetitive DNA silencing is mostly due to polycomb protein group silencing mechanisms. In C. elegans, the SKN-1 gene mediates caloric restriction.
Mice
The pathway in mice shows that the increase in mitochondrial number and activity may work via mitochondrial sirtuins (SIRT3, 4, and 5) or by reducing reactive oxygen species (ROS). Resveratrol and CR may increase SIRT1 activity, which is part of an autoregulatory feedback loop that includes the enzyme endothelial nitric oxide synthase (eNOS). In this aspect, eNOS and SIRT1 have a “reciprocal causation” relationship, where SIRT1 deacetylates eNOS and eNOS activates SIRT1. In mice and probably humans, SIRT1 may increase PGC-1α activity by deacetyation of several lysine side chains, increasing the ability of PGC-1α to co-activate genes.
[Rodgers et al., 2005] and [Gerhart-Hines et al., 2007]). By this mechanism, SIRT1 can increase mitochondrial biogenesis. In humans, CR up regulates SIRT1, eNOS, TFAM, PGC-1 α, and AMPK. In humans, CR down regulates mTOR, the Insulin/IGF-1 pathway, and NF-kB. For these reasons, it is more difficult to give SIRT1 all of the credit for the effects of CR in humans.
The Stress Resistance and Anti-cancer effects of Caloric Restriction and SIRT1
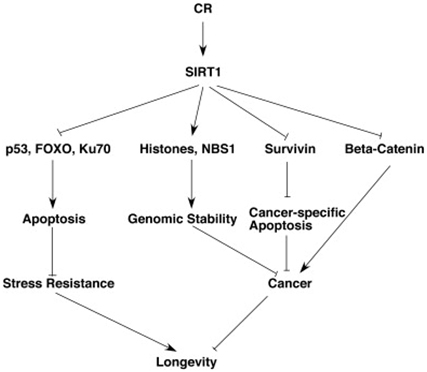
Although this blog is primarily about gene silencing of repetitive DNA, it is important to point out that caloric restriction is one of the most powerful cancer prevention strategies that has been found. CR is even more effective than polyphenol-based chemoprevention in in vitro cell studies and in vivo animal studies. This anti-cancer effect of SIRT1 is mediated via 5 mechanisms:
1. Survivin deacetylation: this induces cancer-specific apoptosis
2. β-catenin deacetylation: this prevents cancer growth
3. Notch deacetylation: this inhibits cancer angiogenesis
4. Histone & NBS1 deacetylation: this promotes genomic stability by gene silencing
5. Nrf2 and FOXO3 deacetylation: this increases gene expression of antioxidant enzymes
The diagram below illustrates this:
SIRT1 increases stress resistance by three separate mechanisms. SIRT1 deacetylates the FOXO transcription factors, thereby activating genes for anti-apoptosis mechanisms. SIRT1 also deacetylates p53, which makes it less likely to induce apoptosis. SIRT1 also activates Ku70, which activates DNA repair mechanisms. As a result of the 6 pathways mentioned above, SIRT1 increases longevity by reducing cancer and increasing stress resistance. Both of these pathways play a role in the longevity effects of SIRT1. Unfortunately, SIRT1 expression declines with aging. As a result, all of these cancer prevention mechanisms and these stress resistant mechanisms decline with aging.
references:
1. Xiaolei Qiu, Katharine V. Brown, Yehu Moran, Danica Chen, Sirtuin regulation in Calorie Restriction, Biochimica et Biophysica Acta (BBA) – Proteins and Proteomics, Volume 1804, Issue 8, August 2010, Pages 1576–1583
2. Hongwei Yao, Irfan Rahman, Perspectives on translational and therapeutic aspects of SIRT1 in inflammaging and senescence, Biochemical Pharmacology, Volume 84, Issue 10, 15 November 2012, Pages 1332–1339
3. Xiaolei Qiu1, 3, Katharine Brown1, 3, Matthew D. Hirschey2, Eric Verdin2, Danica Chen, Calorie Restriction Reduces Oxidative Stress by SIRT3-Mediated SOD2 Activation, Cell Metabolism, Vol 12(6), pp 662-667, December, 2010
The “Silencing Signatures” of SIRT2 and SIRT6
Whereas Sir2 appears to play the role of “gene silencer” in Baker’s yeast, in humans gene silencing is done primarily by SIRT1, SIRT2, SIRT3, and SIRT6. Each of these SIRTs have other functions besides gene silencing, however. They are “redox sensitive” enzymes, like all of the Sirtuins. Since SIRT2 and SIRT6 are two good examples of the two primary “silencing signatures” seen on histones, I will limit the discussion of gene silencing to these two good example Sirtuins.
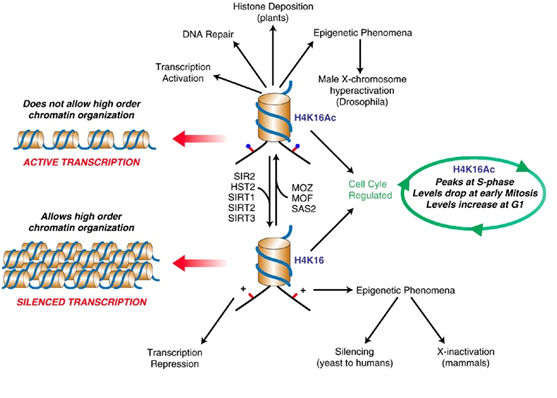
SIRT2 – The gene silencer with the “H4K16 silencing signature”
SIRT2 is a Class III Histone deacetylase. Like SIRT1, SIRT2 has many targets for deacetylation in humans, including FOXO1, p53, FOXO3, alpha-tubulin, and Histones. SIRT2 deacetylates the K16 lysine on the tail of histone protein 4. The acetylation status of lysine 16 on Histone H4 has a critical role in multiple functions in chromatin regulation. Acetylation of H4K16 is associated with active transcription and open chromatin. H4K16 deacetylation is an important silencing mechanism that appears to be “silencing signature” common to SIRT1, SIRT2, and SIRT3. However Here is a diagram illustrating how this works.
Diagram reference: http://www.readcube.com/articles/10.1038/sj.onc.1210617
Other references for text:
Christopher L Brooks, Wei Gu, p53 Activation: A Case against Sir, Cancer Cell, Vol 13(5), pp 377-378, May 6, 2008
Takashi Nakagawa, Leonard Guarente, Urea cycle reguation by mitochondrial sirtuin, SIRT5, Aging, Vol 1(6), pp 578-581, June, 2009.
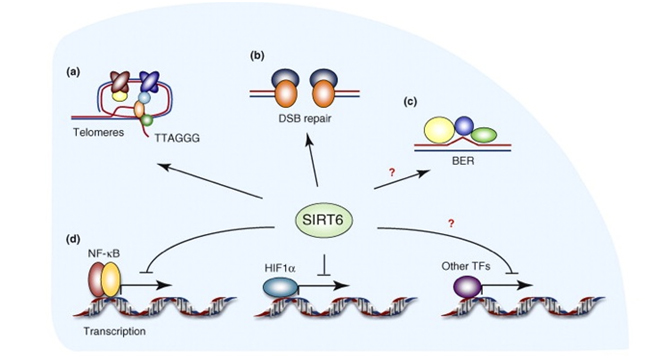
SIRT6 – The ‘Split End Repair” and the “gene silencing signatures” of SIRT6
Whereas SIRT2 was primarily involved with transcription factor deacetylation and H4K16 mediated gene silencing, SIRT6 appears to have multiple roles in maintaining telomere integrity, preventing telomere fusions, and DNA repair. It does have a secondary role in in gene silencing via two other unique “silencing signatures”, but the other functions appear to be more important than gene silencing. Here is a diagram illustrating the main functions of SIRT6.
diagram reference: http://www.sciencedirect.com/science/article/pii/S0968000410001428
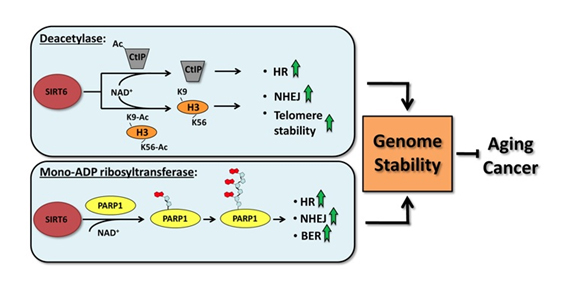
SIRT6 is both a protein deacetylator and a ADP ribosylator. Both functions of the SIRT6 enzyme are very important for genome stability. The diagram below illustrates how this works:
picture reference: http://www.impactaging.com/papers/v3/n9/full/100389.html
As a deacetylator, SIRT6 has three unique “silencing signatures”. SIRT6 deacetylates two specific lysine side chains on the H3 histone tail – the K56 lysine and the K9 lysine. For this reason, H3K9 or H3K56 deacetylation is a tell tail “sign” that SIRT6 has come by. Also, SIRT6 deacetylates CtIP.
The most dangerous event for genomic stability is a double-stranded DNA break. Unrepaired DNA leads to irregular gene expression, cell cycle arrest, apoptosis, and cancer. To ensure that there are two “redundant” systems for DNA repair, two separate repair systems have evolved – homologous recombination (HR) and non-homologous end joining (NHEJ). The first evidence that SIRT6 was involved in double-stranded DNA breaks came from knockout mice experiments. These mice exhibited a very high incidence of chromosomal rearrangements and breaks. They were very sensitive to ionizing radiation, especially gamma radiation. Now there is good evidence that SIRT6 specifically recruited to break sites after DNA damage. SIRT6 functions in NHEJ by stabilizing the NHEJ haloenzyme, DNA-PK at the sites of the double stranded breaks. SIRT6 also promotes HR by deacetylation of the end resection protein, CtIP. Overexpression of SIRT6 stimulates DSB repair through both the HR and NHEJ pathways by 3-fold. SIRT6 does this by its ADP-ribosylation function. SIRT6 ribosylates the upstream DSB repair factor, PARP1, at lysine 521, thereby stimulating its poly-ADP ribosylation activity. PARP1 facilitates the recruitment of the MRN complex to double stranded breaks, plays a role in the activation of ATM and helps to direct the choice between the NHEJ and HR repair pathways. Additionally, PARP1 is required to promote a non-canonical form of NHEJ (Alternate NHEJ pathway). By this mechanism, SIRT6 can stimulate NHEJ in the absence of DNA-PK.
reference: Repairing split ends: SIRT6, mono-ADP ribosylation and DNA repair
Stress Preconditioning before you drink your Wine – evidence for a hormetic ROS dose preconditioning for DNA repair.
An interesting phenomena was found by a group at the University of Rochester recently. They were studying the effects of SIRT6 on DNA Strikingly, when cells were pretreated with oxidative stressors prior to overexpression of SIRT6, DSB repair efficiency was massively stimulated by up to 16-fold, suggesting that SIRT6 specifically integrates stress signaling to prime the DNA repair machinery in response to oxidative stress. Not only did ROS preconditioning increase the intensity of the double-stranded break repair response, it recruited SIRT6 to the site of the double stranded break much sooner (30 minute recruitment time vs 8 hour recruitment time). This means that some stress preconditioning before exposure to DNA damage (for instance, chemotherapy, X-rays, or cigarette smoke) will increase the speed and intensity of the DNA repair response by SIRT6. It appears that that both the deacetylation function and the mono-ADP ribosylation function of SIRT6 is needed for this stress response. It appears that both SIRT6 and SIRT1 localizes to double-stranded DNA breaks in response to DNA damage.
reference: Repairing split ends: SIRT6, mono-ADP ribosylation and DNA repair
POLYCOMB GROUP PROTEIN GENE SILENCING
The second “song of silence” referred to in the title of this blog is via polycomb group proteins which can silence genes in plants, insects and animals. “Polycomb proteins play a key role in developmental gene regulation in most multicellular organisms, from plants to mammals. They were first defined in Drosophila as factors which allow cells to ‘remember’ patterns of gene expression through successive cell divisions, providing a memory of the cell fate or identity established through differentiation and development. Polycomb proteins play a key role in developmental gene regulation in most multicellular organisms, from plants to mammals. They were first defined in Drosophila as factors which allow cells to ‘remember’ patterns of gene expression through successive cell divisions, providing a memory of the cell fate or identity established through differentiation and development. In mammals, polycomb proteins have important roles throughout an organism’s life. In embryonic stem (ES) cells, they play a crucial role in restraining the cells from going down differentiation pathways, targeting around 1500 genes. They are vital in the establishment and maintenance of different cell types. Polycomb proteins are also found on the inactive X chromosome. — Polycomb proteins function in multiprotein complexes which are recruited to and silence selected genes. They accomplish silencing by chemically modifying the histone proteins around which the DNA is entwined. — Polycomb complexes exist in two forms – PRC1 and PRC2. PRC2 loads onto the chromatin first, modifying a lysine on one of the resident histone proteins by adding methylation groups – H3K27me3. This methylated lysine is recognised by a protein called CBX in the PRC1 complex. PRC1 has a different chromatin modifying activity known as ubiquitylation. Bringing PRC1 onto the site leads to ubiquitylation of another histone protein, H2A. It is this modification that inhibits transcription on the genes located at that site, possibly by blocking the enzyme which transcribes the DNA into RNA(ref).” Reference RYBP-PRC1 Complexes Mediate H2A Ubiquitylation at Polycomb Target Sites Independently of PRC2 and H3K27me3.
A number of publications have focused on the gene-silencing actions of polycomb group proteins, for example, Silencing by plant Polycomb-group genes requires dispersed trimethylation of histone H3 at lysine 27, “In plants and animals, Polycomb-group (Pc-G) genes mediate mitotically stable repression of targets such as homeotic genes that are critical for developmental patterning and growth control. An antagonistic group of proteins, the trithorax-group (trx-G), act as activators. Both groups act to maintain on/off patterns of transcription defined early in development, rather than to set up these patterns, and are important for maintaining cell fates. Although the Pc-G and trx-G have long been thought to cause epigenetic changes in chromatin structure, due to the stable but ultimately reversible alterations in gene activity they promote, the mechanistic basis for their activity has been mysterious. Recent studies have implicated histone modifications as an important component of epigenetic changes. A variety of modifications on the amino-tails of histones have been characterised, of which methylation of lysine residues has been thought to be particularly stable (Jenuwein and Allis, 2001; Fischle et al, 2003). The consequence of lysine methylation can differ both according to which lysine residue is modified and also as to how many methyl groups are added—lysine residues can be mono-, di- or trimethylated (Bannister et al, 2002). For example, methylation of histone H3 at lysine 9 (H3K9) or lysine 27 (H3K27) is generally correlated with transcriptional repression, whereas methylation at lysine 4 (H3K4) is predominantly associated with transcriptional activity (Jenuwein and Allis, 2001; Peters et al, 2003; Ringrose and Paro, 2004). In addition, the level of methylation is important, for example H3K9me3 shows a different distribution from H3K9me1 and H3K9me2 in mammals (Peters et al, 2003). Several enzymes with histone lysine demethylase activity have now been identified, indicating that methylation can be rapidly reversed (Shi et al, 2004; Tsukada et al, 2006).”
Many publications focus on the roles of polycomb group proteins in cancer. For example, Polycomb group protein gene silencing, non-coding RNA, stem cells, and cancer (2011). “Epigenetic programming is an important facet of biology, controlling gene expression patterns and the choice between developmental pathways. The Polycomb group proteins (PcGs) silence gene expression, allowing cells to both acquire and maintain identity. PcG silencing is important for stemness, X chromosome inactivation (XCI), genomic imprinting, and the abnormally silenced genes in cancers. Stem and cancer cells commonly share gene expression patterns, regulatory mechanisms, and signalling pathways. Many microRNA species have oncogenic or tumor suppressor activity, and disruptions in these networks are common in cancer; however, long non-coding (nc)RNA species are also important. Many of these directly guide PcG deposition and gene silencing at the HOX locus, during XCI, and in examples of genomic imprinting. Since inappropriate HOX expression and loss of genomic imprinting are hallmarks of cancer, disruption of long ncRNA-mediated PcG silencing likely has a role in oncogenesis. Aberrant silencing of coding and non-coding loci is critical for both the genesis and progression of cancers. In addition, PcGs are commonly abnormally overexpressed years prior to cancer pathology, making early PcG targeted therapy an option to reverse tumor formation, someday replacing the blunt instrument of eradication in the cancer therapy arsenal.”
Other recent publications relating PcG silencing to cancer include:
Gene silencing by the Polycomb group proteins and associations with cancer
Polycomb silencers control cell fate, development and cancer
Polycomb group proteins: navigators of lineage pathways led astray in cancer
Cell Death and Disease – Roles of the Polycomb group proteins in stem cells and cancer
There is a significant literature related to polycomb group silencing. Topics include signaling in cancers, signaling in plant, insect and human development, and details of the silencing process. 2013 publications include:
Polycomb silencing of the Drosophila 4E-BP gene regulates imaginal disc cell growth.
Epigenetic control of female puberty.
DNA methylation in the malignant transformation of meningiomas.
KDM2B promotes pancreatic cancer via Polycomb-dependent and -independent transcriptional programs.
CCN3/NOV gene expression in human prostate cancer is directly suppressed by the androgen receptor.
Genomic and gene-level distribution of histone H3 dimethyl lysine-27 (H3K27me2) in Arabidopsis.
A gene loop containing the floral repressor FLC is disrupted in the early phase of vernalization.
[miRNA in HTLV-1 related disease].
Site-specific silencing of regulatory elements as a mechanism of X inactivation
DMNT3 GENE SILENCING
The third “song of silence” referred to in this blog title in epigenetic gene silencing brought about by DNMT3 methyltransferases. This subject has already been discussed in detail in the previous blog entry PART 2: Slaying Two Dragons with One Hail of Stones: The Silencing Of Good Genes In Aging And Cancer. “Gene silencing by epigenetic mechanisms is an ordered series of events that normally starts with the methylation of cytosine residues. Scientists studying gene silencing have observed two distinct kinds of CpG methylation: one that is age-related that occurs in all tissues as a function of aging, and a cancer-related pattern of methylation that only occurs in cancerous cells. It is important to point out that with aging, there is a global loss of methylation at sites other than CpG islands. The hypermethylation of CpG islands is the exception to this universal decline in DNA methylation with aging. This can be explained in part by the fact that there are different DNA methyltransferases for these different regions of the genome. DNMT1 is responsible for the maintenance of hemimethylated DNA, whereas DNMT3a and 3b are responsible for the methylation of unmethylated DNA. For this reason, it appears that DNMT3a and 3b are responsible for gene silencing.”
PART B
A master list of drugs and natural compounds for cancer chemoprevention
You may download such a list as a Word document by clicking on link below.
Note on bioavailability of plant polyphenols and nano-particle delivery
This blog post touches on biological activities and health benefits of certain plant polyphenols (e.g. Resveratrol) as do a great many other postings over the years. It should be pointed out that for many plant polyphenols, biological impacts observed in vitro may not be fully realizable in vivo based on normal oral ingestion. There are several reasons for that. 1. Chemical structures can be altered by gut bacteria. Sometimes the gut biome helps us in this regard, an example being gut actions on the polyphenols in chocolate (protoanthocyanadins). But in states of altered gut micro biome, we can’t always count on the bacteria. 2. Poor absorption. This is a major issue with many of the polyphenols, especially curcumin. 3. Metabolism by the liver via the first pass effect, this destroys a lot of the polyphenols. The metabolites are not as good signaling compounds. 4. Glycosylation. This makes substances more water soluble, but then they are secreted faster. Also, glycosylation reduces the ability of a substance to become part of the mitochondrial membrane and work as a hydrophobic mitochondrial specific antioxidant. (I am now convinced that the chloroplast outer membranes are places where stress-defensive phytochemicals migrate to in plant cells. In us humans on the other hand, the mitochondrial membranes are where we can control excess baseline ROS leak that occurs in old mitochondria.) 5. Last of all, plant polyphenols tend to be very prone to oxidation. This means they are inactivated with heat (instantly) in water (4 hrs), and in the presence of oxygen, especially with cation catalysts like unliganded iron or copper.
Indeed, poor bioavailability is probably the major reason why clinical trials of certain phyto- substances like resveratrol and curcumin against disease indications like cancers and Alzheimer’s disease have not produced positive results.
We believe there is an elegant solution on the horizon to this issue of poor bioavailability of health-producing phyto substances. That is, the use of nanoparticle polyphenols. Very recently, there has been significant research related to nano-particle delivery of drugs and phyto substances. Because of the importance of the bioavailability issue and the emerging importance of nano particle delivery, we plan to produce a blog on the bioavailability issue and on nano particle delivery in the near future.
DATA DISCLAIMER:
The tables and compilations of data in this and the other blog entries in the Two Dragons series are intended to be illustrative of the main points made in the blog entries. The data are compiled from various sources, in most cases are incomplete, and may contain occasional errors.
MEDICAL DISCLAIMER
FROM TIME TO TIME, THIS BLOG DISCUSSES DISEASE PROCESSES. THE INTENTION OF THOSE DISCUSSIONS IS TO CONVEY CURRENT RESEARCH FINDINGS AND OPINIONS, NOT TO GIVE MEDICAL ADVICE. THE INFORMATION IN POSTS IN THIS BLOG IS NOT A SUBSTITUTE FOR A LICENSED PHYSICIAN’S MEDICAL ADVICE. IF ANY ADVICE, OPINIONS, OR INSTRUCTIONS HEREIN CONFLICT WITH THAT OF A TREATING LICENSED PHYSICIAN, DEFER TO THE OPINION OF THE PHYSICIAN. THIS INFORMATION IS INTENDED FOR PEOPLE IN GOOD HEALTH. IT IS THE READER’S RESPONSIBILITY TO KNOW HIS OR HER MEDICAL HISTORY AND ENSURE THAT ACTIONS OR SUPPLEMENTS HE OR SHE TAKES DO NOT CREATE AN ADVERSE REACTION






fullerene C60 results in increased expression of SIRT6 mRNA – http://rgd.mcw.edu/rgdweb/report/annotation/table.html?term=CHEBI:33128&id=1305216
(see the supplemental table in the mmc1.xls attachment for PMID 19167457)
Pingback: PART 2: Slaying Two Dragons with One Hail of Stones: The Silencing Of Good Genes In Aging And Cancer – And How Polyphenols Can Prevent That | AGING SCIENCES – Anti-Aging Firewalls
http://www.ncbi.nlm.nih.gov/pubmed/23636469 – Suppression of native defense mechanisms, SIRT1 and PPARγ, by dietary glycoxidants precedes disease in adult humans; relevance to lifestyle-engendered chronic diseases
jz99
First of all, thanks for your recent stream of interesting comments and citations in this blog. I have found everything you have cited to be both interesting and relevant.
With regard to your post on dietary glycoxidants, this seems to be a “full circle”finding. That is, years ago AGEs and glycation were thought to be very important phenomena in aging, but in recent years they seem to have been crowded out into near oblivion by a host of current issues related to molecular biology, epigenetic’s, and stem cells. I listed Tissue Glycation as one of the 14 basics theories of aging when I first wrote my treatise on aging (at http://www.vincegiuliano.name/Antiagingfirewalls.htm) some six years ago. The publication you cite directly relates dietary AGEs to suppression of stress defense mechanisms such as SIRT1 and PPARγ and goes a long way towards explaining why AGEs have some of the effects we have long known them to have.
With regard to C-60 fullerenes increasing the expression of SIRT6 mRNA, I am not too surprised although I believe what these fullerenes do is still only partially understood at best. Perhaps this partially explains the purported life-extending properties of buckyballs. As mentioned in another comment, I hope to come back to writing about buckyballs and possibly other nano-substances in another blog entry soon.
Vince
Hi,
a relevant practical question is the appropiate way to administrate Sirt activators like Resveratrol/Quercetin. Is it best to use it in a day to day basis, or a big intake once per week in a fasting day?. Later approach could enhance the benefits of fasting and sirt activators intake.
Best.
Pingback: THE HORMESIS BARS | AGING SCIENCES – Anti-Aging Firewalls
Pingback: Quorum sensing Part 1: quorum sensing inhibition via phytochemicals – a new approach against infectious diseases. | AGING SCIENCES – Anti-Aging Firewalls
Pingback: Silencing, Heterochromatin and DNA Double Strand Break Repair · WWW.DBESTREVIEW.COM