Certain plant polyphenols not only exercise general positive health effects but also inhibit oncogenic transformation or the proliferation of cancer cells. Or, they outright kill cancer cells. Included are a substantial number of cancer-fighting plant polyphenols, some of which are found in familiar substances like green tea, turmeric, ginger, garlic, broccoli and bitter melon. This blog entry is about the biological pathways and epigenetic transformations through which these cancer-fighting effects take place. I start with short descriptions of six semi-independent modes of epigenetic action through which certain polyphenols attack cancer cells – the six knockout punches delivered by these polyphenols. Then I cite a selection of research publications supporting this general picture and filling in the details.
Picture source Image source
Cancer-fighting polyphenols (CFPs) exert their actions through at least six mechanisms:
First knockout punch mechanism – downregulation of expression of NF-kappaB
CFPs downregulate the expression of the pro-inflammatory intermediary NF-kappaB. Cancer cells proliferate most readily in a pro-inflammatory environment and find anti-inflammatory environments hostile.
Second knockout punch mechanism – CFPs creates stress and stress signaling in cells
In normal cells, the stress generated by normal dietary intake of CFPs is within the hormetic range of response. Nrf2 translocates into the nucleus, ARE genes are activated including anti-oxidant and Phase 2 detoxifying genes. The net result is health-producing for the cell. The same is true when the stress is due to normal doses of CFP supplements. In cancer cells which are relatively deregulated to start with, the stress generated by CFP intake can be beyond the hormetic range, with the result being damage or death to the cancer cell.
Third knockout punch mechanism – HDAC inhibition which re-activates tumor suppressor genes.
CFPs act as HDAC inhibitors or HATs (histone deacetylase inhibitors or histone acetyl transferases). Expression of protective pro-apoptotic genes like P53 and P21 is turned on in acetylated cancer cells while it is usually turned off in deacetylated cancer cells. So, sensing self-damage, acetylated cancer cells are more likely to terminate themselves via apoptotic death.
Fourth knockout punch mechanism – impact on DNA repair machinery
For the same reasons related to both non-hormetic oxidative stress and histone acetylation, cancer cells in an acetylated state are more likely to experience damage to their DNA repair machinery.
Fifth knockout punch – impact of CFPs on mRNAs
CFPs act on mRNAs so as to induce tumor suppressor mRNAs and block oncogenic mRNAs
Sixth knockout punch mechanism – – CFPs inhibit DNA methyltransferases (DNMT) in cancer cells, thus blocking the methylation and inactivation of tumor suppressor genes in cancers.
These mechanisms are further elucidated in the research publications cited below.
What are the major CFPs?
A listing of many of them is found in the abstract for the publication Molecular targets of dietary agents for prevention and therapy of cancer, although in 2006, the time of that publication, the epigenetic mechanisms were only partially understood. “While fruits and vegetables are recommended for prevention of cancer and other diseases, their active ingredients (at the molecular level) and their mechanisms of action less well understood. Extensive research during the last half century has identified various molecular targets that can potentially be used not only for the prevention of cancer but also for treatment. However, lack of success with targeted monotherapy resulting from bypass mechanisms has forced researchers to employ either combination therapy or agents that interfere with multiple cell-signaling pathways. In this review, we present evidence that numerous agents identified from fruits and vegetables can interfere with several cell-signaling pathways. The agents include curcumin (turmeric), resveratrol (red grapes, peanuts and berries), genistein (soybean), diallyl sulfide (allium), S-allyl cysteine (allium), allicin (garlic), lycopene (tomato), capsaicin (red chilli), diosgenin (fenugreek), 6-gingerol (ginger), ellagic acid (pomegranate), ursolic acid (apple, pears, prunes), silymarin (milk thistle), anethol (anise, camphor, and fennel), catechins (green tea), eugenol (cloves), indole-3-carbinol (cruciferous vegetables), limonene (citrus fruits), beta carotene (carrots), and dietary fiber. For instance, the cell-signaling pathways inhibited by curcumin alone include NF-kappaB, AP-1, STAT3, Akt, Bcl-2, Bcl-X(L), caspases, PARP, IKK, EGFR, HER2, JNK, MAPK, COX2, and 5-LOX. The active principle identified in fruit and vegetables and the molecular targets modulated may be the basis for how these dietary agents not only prevent but also treat cancer and other diseases. This work reaffirms what Hippocrates said 25 centuries ago, let food be thy medicine and medicine be thy food.”
The overall epigenetic actions of CFPs on regulating cancers is summarized in the Sept 2012 publiction Cancer Chemoprevention and Nutri-Epigenetics: State of the Art and Future Challenges: “The term “epigenetics” refers to modifications in gene expression caused by heritable, but potentially reversible, changes in DNA methylation and chromatin structure. Epigenetic alterations have been identified as promising new targets for cancer prevention strategies as they occur early during carcinogenesis and represent potentially initiating events for cancer development. Over the past few years, nutri-epigenetics – the influence of dietary components on mechanisms influencing the epigenome – has emerged as an exciting new field in current epigenetic research. During carcinogenesis, major cellular functions and pathways, including drug metabolism, cell cycle regulation, potential to repair DNA damage or to induce apoptosis, response to inflammatory stimuli, cell signalling, and cell growth control and differentiation become deregulated. Recent evidence now indicates that epigenetic alterations contribute to these cellular defects, for example epigenetic silencing of detoxifying enzymes, tumor suppressor genes, cell cycle regulators, apoptosis-inducing and DNA repair genes, nuclear receptors, signal transducers and transcription factors by promoter methylation, and modifications of histones and non-histone proteins such as p53, NF-κB, and the chaperone HSP90 by acetylation or methylation.The present review will summarize the potential of natural chemopreventive agents to counteract these cancer-related epigenetic alterations by influencing the activity or expression of DNA methyltransferases and histone modifying enzymes. Chemopreventive agents that target the epigenome include micronutrients (folate, retinoic acid, and selenium compounds), butyrate, polyphenols from green tea, apples, coffee, black raspberries, and other dietary sources, genistein and soy isoflavones, curcumin, resveratrol, dihydrocoumarin, nordihydroguaiaretic acid (NDGA), lycopene, anacardic acid, garcinol, constituents of Allium species and cruciferous vegetables, including indol-3-carbinol (I3C), diindolylmethane (DIM), sulforaphane, phenylethyl isothiocyanate (PEITC), phenylhexyl isothiocyanate (PHI), diallyldisulfide (DADS) and its metabolite allyl mercaptan (AM), cambinol, and relatively unexplored modulators of histone lysine methylation (chaetocin, polyamine analogs). So far, data are still mainly derived from in vitro investigations, and results of animal models or human intervention studies are limited that demonstrate the functional relevance of epigenetic mechanisms for health promoting or cancer preventive efficacy of natural products. Also, most studies have focused on single candidate genes or mechanisms research has the potential to explore nutri-epigenomics at a genome-wide level to understand better the importance of epigenetic mechanisms for gene regulation in cancer chemoprevention.”. With the emergence of novel technologies such as next-generation sequencing, future
Background for first knockout punch mechanism – downregulation of expression of NF-kappaB
Regarding cancer-fighting polyphenols: I have discussed the anti-cancer activities of CFPs in a number of blog entries. In the blog entry New, emerging and potential treatments for cancers: Part 3 – selected less-known phytochemicals that have long been used in traditional Chinese medicine – focus on gambogic and gambogenic acids I related “ Many traditional Chinese medicines have been extensively studied in China during the last 10-20 years using the current tools and intellectual frameworks of modern Western science. These medicines have been looked at in terms of their detailed chemical structures, their proteomic properties, the molecular biological pathways through which they work, their gene activation and epigenetic properties, their pharmacological properties, etc, This work has generally been of high quality and has resulted in thousands or tens of thousands of research reports, many of them published in highly respected Western journals. Abstracts to these publications can be found in the definitive US National Library of Medicine database pubmed.org. – A good place to start is with the July 2011 e-publication Anti-cancer natural products isolated from chinese medicinal herbs, as to be expected written by a team of Chinese researchers. “In recent years, a number of natural products isolated from Chinese herbs have been found to inhibit proliferation, induce apoptosis, suppress angiogenesis, retard metastasis and enhance chemotherapy, exhibiting anti-cancer potential both in vitro and in vivo. This article summarizes recent advances in in vitro and in vivo research on the anti-cancer effects and related mechanisms of some promising natural products. These natural products are also reviewed for their therapeutic potentials, including flavonoids (gambogic acid, curcumin, wogonin and silibinin), alkaloids (berberine), terpenes (artemisinin, β-elemene, oridonin, triptolide, and ursolic acid), quinones (shikonin and emodin) and saponins (ginsenoside Rg3), which are isolated from Chinese medicinal herbs. In particular, the discovery of the new use of artemisinin derivatives as excellent anti-cancer drugs is also reviewed.”
Regarding expression of NF-kappaB: From my Treatise: “An important line of epigenomic research relates to NF-kappaB signaling. NF-kappaB is a nuclear transcription factor involved in cell signaling, i.e. a protein that binds to a specific sequence of DNA. It is present in a latent (non-activated) form in many cell types. On the one hand, expression of NF-kB appears to be one of the body’s regulatory means for handling situations of stress, cancer, damage or disease. In eukaryotic cells NF-kB is an important regulator of genes that control cell proliferation and cell survival. NF-kB regulates anti-apoptotic genes that protect healthy cells from cell death and activates the expression of genes that keep cells proliferating. On the other hand, activated NF-kB binding to genes has long been known to play a central role in promoting runaway inflammation and inflammation’s negative consequences. — These consequences include promotion of angiogenesis, proliferation, metastasis and invasiveness in cancer tumors, autoimmune diseases, neurodegenerative diseases and contributing to the activation of human immunodeficiency virus (HIV) leading to AIDS. — There appears to be increasing evidence that inhibition of expression of NF-kB could be a key approach for fighting cancers, controlling inflammatory diseases, AIDS, neurodegenerative conditions like Parkinson’s Disease and a number of other significant age-related maladies. — Recent studies position NF-kB even more centrally with respect to longevity. It is likely that NF-kB expression is central to a programmed set of changes which we call aging. One study(ref1) confirms that that in multiple mammalian tissues (including skin fibroblasts, kidney, cortex, kidney medulla, abdominal muscle, skeletal muscle, and brain), aging involves continuing changes in expression of hundreds of genes. And, further, NF-kB signaling appears to be a major regulator of gene expression related to the aging progress. In fact, by inhibiting NF-kB cell signaling the researchers were able to cause the epidermal tissue of old mice to revert to the state of very young mouse tissue, both in observable characteristics and in genetic expression profile. The authors show that NF-kB cell signaling is a meta-factor for determining aging of nine other key cell types as well, and they argue that the results should apply equally to humans and other mammals.” The relationship of NF-kappaB expression to oncogenic processes. cancer proliferation and metastasis runs deep and wide. See for example The activated NF-kappaB-Snail-RKIP circuitry in cancer regulates both the metastatic cascade and resistance to apoptosis by cytotoxic drugs.
Regarding CFPs and the expression on NF-kappaB: Discussion in my treatise as well as many blog entries point to how CFPs inhibit the expression of NF-kappaB.
“Regarding proinflammatory environments and cancer susceptibility: It has long been known that a pro-inflammatory environment can significantly enhance cancer susceptibilities. The 2002 publication Inflammation and cancer pointed out: “Recent data have expanded the concept that inflammation is a critical component of tumour progression. Many cancers arise from sites of infection, chronic irritation and inflammation. It is now becoming clear that the tumour microenvironment, which is largely orchestrated by inflammatory cells, is an indispensable participant in the neoplastic process, fostering proliferation, survival and migration. In addition, tumour cells have co-opted some of the signalling molecules of the innate immune system, such as selectins, chemokines and their receptors for invasion, migration and metastasis. These insights are fostering new anti-inflammatory therapeutic approaches to cancer development. — The functional relationship between inflammation and cancer is not new. In 1863, Virchow hypothesized that the origin of cancer was at sites of chronic inflammation, in part based on his hypothesis that some classes of irritants, together with the tissue injury and ensuing inflammation they cause, enhance cell proliferation1. Although it is now clear that proliferation of cells alone does not cause cancer, sustained cell proliferation in an environment rich in inflammatory cells, growth factors, activated stroma, and DNA-damage-promoting agents, certainly potentiates and/or promotes neoplastic risk.” Inflammation ,au contribute to cancers via engendering higher rate of DNA mutations. An author of a 2011 study reported “”Our study shows that miR-155 is upregulated by inflammatory stimuli and that overexpression of miR-155 increases the spontaneous mutation rate, which can contribute to tumorigenesis.”
Together, points a. – d. above characterize the first mechanism through which CFPs combat cancers: CFPs downregulate the expression of the pro-inflammatory intermediary NF-kappaB. Cancer cells proliferate most readily in a pro-inflammatory environment and find anti-inflammatory environments hostile.
Background for the second knockout punch mechanism – CFPs creates stress and stress signaling in cells
In normal cells, the stress associated with CFP intake is within the hormetic range of response, Nrf2 translocates into the nucleus, ARE genes are activated including anti-oxidant and Phase 2 detoxifying genes. The net result is health-producing for the cell. See the blog entries Nrf2: Part 1 – a new view on the control of oxidative damage and generation of hormetic effects. The pivotal role of Nrf2. Part 3 – Part 3 – Is promotion of Nrf2 expression a viable strategy for human human healthspan and lifespan extension?, and Part 2 – foods, phyto-substances and other substances that turn on Nrf2.
However, in cancer cells which are relatively deregulated to start with, the stress created by ingestion of CFPs can be beyond the hormetic range, with the result being damage or death to the cancer cell. See the blog entries Mitohormesis and Radiation Hormesis for discussion of hormetic range. For example, for human head and neck squamous cell carcinoma cells, DNA damage has been shown to be caused by resveratrol(ref). Grape seed extract likewise causes DNA damage to squamous cell carcinoma cells (ref). Grape seed extract leads to apoptotic death of human prostate carcinoma DU145 cells via caspases activation accompanied by dissipation of mitochondrial membrane potential and cytochrome c release(ref). Sulphoraphane, a naturally occurring isothiocyanate induces apoptosis in breast cancer cells by targeting heat shock proteins(ref).
Background for the third knockout punch mechanism – HDAC inhibition which re-activates tumor suppressor genes
Cancer cells tend to be epigenetically dysregulated. This can be seen in several epigenetic dimensions, for example that of gene methylation profiles. The just-published (October 2012) publication LRpath analysis reveals common pathways dysregulated via DNA methylation across cancer types reports: “BACKGROUND: The relative contribution of epigenetic mechanisms to carcinogenesis is not well understood, including the extent to which epigenetic dysregulation and somatic mutations target similar genes and pathways. We hypothesize that during carcinogenesis, certain pathways or biological gene sets are commonly dysregulated via DNA methylation across cancer types. The ability of our logistic regression-based gene set enrichment method to implicate important biological pathways in high-throughput data is well established. RESULTS: We developed a web-based gene set enrichment application called LRpath with clustering functionality that allows for identification and comparison of pathway signatures across multiple studies. Here, we employed LRpath analysis to unravel the commonly altered pathways and other gene sets across ten cancer studies employing DNA methylation data profiled with the Illumina HumanMethylation27 BeadChip. We observed a surprising level of concordance in differential methylation across multiple cancer types. For example, among commonly hypomethylated groups, we identified immune-related functions, peptidase activity, and epidermis/keratinocyte development and differentiation. Commonly hypermethylated groups included homeobox and other DNA-binding genes, nervous system and embryonic development, and voltage-gated potassium channels. For many gene sets, we observed significant overlap in the specific subset of differentially methylated genes. Interestingly, fewer DNA repair genes were differentially methylated than expected by chance. CONCLUSIONS: Clustering analysis performed with LRpath revealed tightly clustered concepts enriched for differential methylation. Several well-known cancer-related pathways were significantly affected, while others were depleted in differential methylation. We conclude that DNA methylation changes in cancer tend to target a subset of the known cancer pathways affected by genetic aberrations.”
Histone deacetylase inhibition (HDACi) is an important emerging approach to cancer therapy.
I have made this point before(ref). HDAC inhibitors are of interest for treating many medical conditions besides cancers including targeting Alzheimer’s disease(ref), other neurodegenerative conditions(ref), and diabetes(ref). While many substances are HDAC inhibitors including a number of CFPs, the drug panobinostat is being particularly studied and experimentally used in the cancer research community. “Panobinostat (LBH-589) is an experimental drug developed by Novartis for the treatment of various cancers. It is a hydroxamic acid[1] and acts as a non-selective histone deacetylase inhibitor (HDAC inhibitor) — Panobinostat inhibits multiple histone deacetylase enzymes, a mechanism leading to apoptosis of malignant cells via multiple pathways.[1] (ref).[2]”
Going back to a 2004 publication Role of histone deacetylase inhibitors in the treatment of cancer (Review): “Acetylation and deacetylation of nucleosomal histones play an important role in the modulation of chromatin structure, chromatin function and in the regulation of gene expression. Histone acetyltransferases (HATs) and histone deacetylases (HDACs) are two opposing classes of enzymes, which tightly control the equilibrium of histone acetylation. An imbalance in the equilibrium of histone acetylation has been associated with carcinogenesis and cancer progression. So far, a number of structurally distinct classes of compounds have been identified as HDAC inhibitors including the short-chain fatty acids, hydroxamates, cyclic tetrapeptides and benzamides. These compounds lead to an accumulation of acetylated histone proteins both in tumor cells and in normal tissues. HDAC inhibitors are able to activate differentiation, to arrest the cell cycle in G1 and/or G2, and to induce apoptosis in transformed or cancer cells. Attention is currently being drawn to molecular mechanisms involving histone deacetylases. An induction of p21(WAF/CIP1) and a suppression of angiogenic stimulating factors have been observed in tumor cells following exposure to HDAC inhibitors. In xenograft models, several HDAC inhibitors have demonstrated antitumor activity with only few side effects. Several clinical trials showed that HDAC inhibitors in well tolerated doses have significant antitumoral activities. A combination of HDAC inhibitors with differentiation-inducing agents and cytotoxic drugs is an innovative therapeutic strategy that carries the potential for significant improvements in the treatment of cancer.”
The 2010 publication Histone deacetylase inhibitors: mechanisms and clinical significance in cancer: HDAC inhibitor-induced apoptosis tells a part of the story. “Epigenic modifications, mainly DNA methylation and acetylation, are recognized as the main mechanisms contributing to the malignant phenotype. Acetylation and deacetylation are catalyzed by specific enzymes, histone acetyltransferases (HATs) and histone deacetylases (HDACs), respectively. While histones represent a primary target for the physiological function of HDACs, the antitumor effect of HDAC inhibitors might also be attributed to transcription-independent mechanisms by modulating the acetylation status of a series of non-histone proteins. HDAC inhibitors may act through the transcriptional reactivation of dormant tumor suppressor genes. They also modulate expression of several other genes related to cell cycle, apoptosis, and angiogenesis. Several HDAC inhibitors are currently in clinical trials both for solid and hematologic malignancies. Thus, HDAC inhibitors, in combination with DNA-demethylating agents, chemopreventive, or classical chemotherapeutic drugs, could be promising candidates for cancer therapy.”
HDAC-inhibiting drugs have been approved for cancer indications by the FDA and are in clinical use. A list of HDAC-inhibiting drugs for cancers and other disease indications in clinical trials in 2010 can be found here. As reported in the 2011 publication Rational therapeutic combinations with histone deacetylase inhibitors for the treatment of cancer, HDAC-inhibiting drugs have been approved by the FDA and are used generally in combination with other drugs. ”Histone deacetylases (HDACs) regulate the acetylation of a variety of histone and nonhistone proteins, controlling the transcription and regulation of genes involved in cell cycle control, proliferation, survival, DNA repair and differentiation. Unsurprisingly, HDAC expression is frequently altered in hematologic and solid tumor malignancies. Two HDAC inhibitors (vorinostat and romidepsin) have been approved by the US FDA for the treatment of cutaneous T-cell lymphoma. As single agents, treatment with HDAC inhibitors has demonstrated limited clinical benefit for patients with solid tumors, prompting the investigation of novel treatment combinations with other cancer therapeutics. In this article, the rationales and clinical progress of several combinations with HDAC inhibitors are presented, including DNA-damaging chemotherapeutic agents, radiotherapy, hormonal therapies, DNA methyltransferase inhibitors and various small-molecule inhibitors. The future application of HDAC inhibitors as a treatment for cancer is discussed, examining current hurdles to overcome before realizing the potential of this new approach.”
Going beyond the above, it is possible now to surface deeper mechanisms through which HDAC inhibitors fight cancers
There are at least two mechanisms through which HDAC inhibitors fight cancers: 1. turning on pro-apoptotic tumor suppresssor genes like P53 and P21 in cancer cells and 2. downregulates the cancer cell’s DNA repair machinery.
It has long been known that histone deacytlases negatively regulate tumor suppressor genes thus promoting carcinogenesis(ref). Similarly HDAC inhibitors can cause re-expression of tumor suppressor genes like P53 which are turned off in cancer cells. For example, benzyl isothiocyanate (BITC), a constituent of edible cruciferous vegetables, decreases viability of cancer cells by causing apoptosis. Its mode of operation appears to involve upregulation of P53(ref), likely to be due to HDAC inhibitory activity.
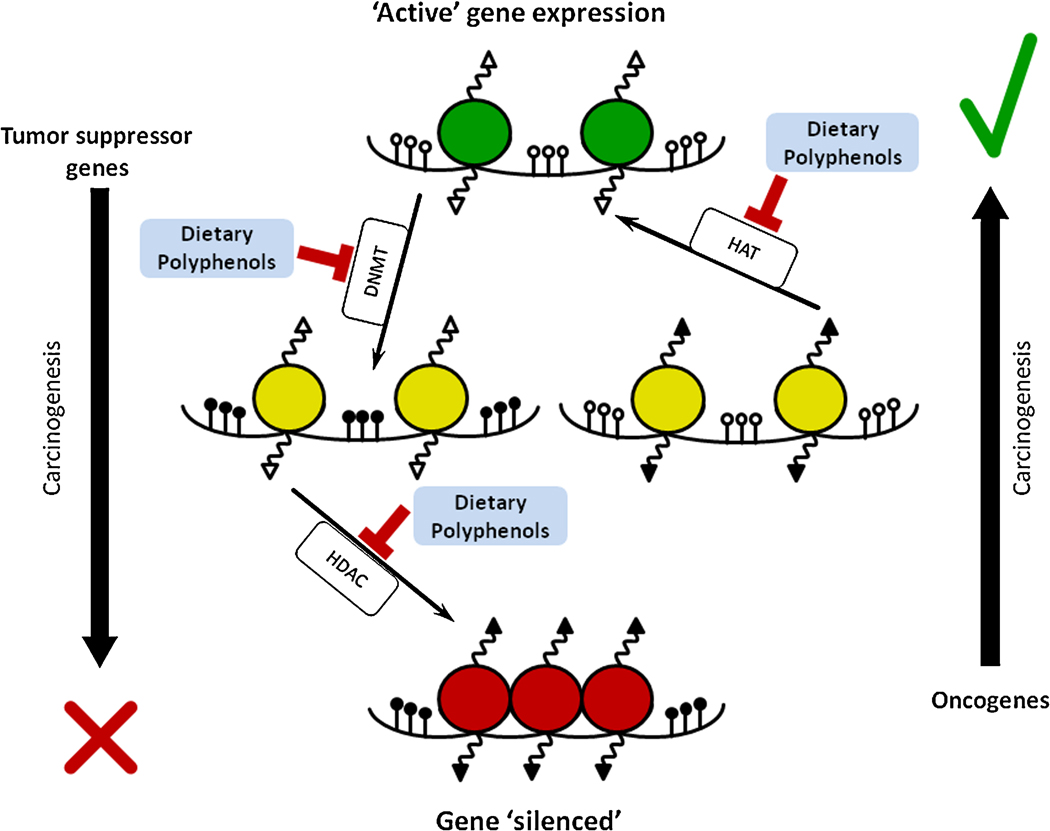
Image source “Effects of dietary polyphenols on the DNA methylation and histone modifications – Simplified scheme demonstrates a number of epigenetic changes that occur during carcinogenesis. In cancers, tumor suppressor genes become “inactivated” (shown as red circles) while oncogenes are “activated” (green circles). Epigenetic gene expression regulation is a complex process and several key enzymes play crucial roles. DNA methyltransferase (DNMT) is responsible for transfer of methyl group to 5′-cytosine. Histone acetylases (HAT) and histone deacetylases (HDAC) are responsible for the acetylation and de-acetylation of lysine residues within histone tails, respectively. Because of these histone modifications, conformational changes in chromatin structure lead to changes in DNA accessibility for transcription regulators and polymerases. Polyphenols can impact these enzymes in specific ways induces reversibility of epigenetic dysregulation in cancer cells(ref).”
The 2010 publication Downregulation of Homologous Recombination DNA Repair Genes by HDAC Inhibition in Prostate Cancer Is Mediated through the E2F1 Transcription Factor discusses the second mechanism in the case of prostate cancer: “Background Histone deacetylase inhibitors (HDACis) re-express silenced tumor suppressor genes and are currently undergoing clinical trials. Although HDACis have been known to induce gene expression, an equal number of genes are downregulated upon HDAC inhibition. The mechanism behind this downregulation remains unclear. Here we provide evidence that several DNA repair genes are downregulated by HDAC inhibition and provide a mechanism involving the E2F1 transcription factor in the process. Methodology/Principal Findings Applying Analysis of Functional Annotation (AFA) on microarray data of prostate cancer cells treated with HDACis, we found a number of genes of the DNA damage response and repair pathways are downregulated by HDACis. AFA revealed enrichment of homologous recombination (HR) DNA repair genes of the BRCA1 pathway, as well as genes regulated by the E2F1 transcription factor. Prostate cancer cells demonstrated a decreased DNA repair capacity and an increased sensitization to chemical- and radio-DNA damaging agents upon HDAC inhibition. Recruitment of key HR repair proteins to the site of DNA damage, as well as HR repair capacity was compromised upon HDACi treatment. Based on our AFA data, we hypothesized that the E2F transcription factors may play a role in the downregulation of key repair genes upon HDAC inhibition in prostate cancer cells. ChIP analysis and luciferase assays reveal that the downregulation of key repair genes is mediated through decreased recruitment of the E2F1 transcription factor and not through active repression by repressive E2Fs. Conclusions/Significance Our study indicates that several genes in the DNA repair pathway are affected upon HDAC inhibition. Downregulation of the repair genes is on account of a decrease in amount and promoter recruitment of the E2F1 transcription factor. Since HDAC inhibition affects several pathways that could potentially have an impact on DNA repair, compromised DNA repair upon HDAC inhibition could also be attributed to several other pathways besides the ones investigated in this study. However, our study does provide insights into the mechanism that governs downregulation of HR DNA repair genes upon HDAC inhibition, which can lead to rationale usage of HDACis in the clinics.




Hi Vincent,
What supplements/herbs do you recommend to take as anti aging? I am an 33 years old man.
Thanks!
Hi Danutz
“Recommend” is too strong a word given the uncertainties associated with biology; I prefer “suggest.” Having said that:
1. A short list of my “top 10 supplements” suitable for a younger person like yourself in good health can be found at http://www.vincegiuliano.name/10topsupplements.htm
I put this list together in 2009 and it still looks pretty good to me.
2. A much more comprehensive list perhaps suitable for an old geeze like me in good health is in my treatise ANTI-AGING FIREWALLS – THE SCIENCE AND TECHNOLOGY OF LONGEVITY at http://www.vincegiuliano.name/Antiagingfirewalls.htm Look for the Table entitled TABLE I – SUPPLEMENTS IN COMBINED FIREWALLS.
Vince
Pingback: PART 1: Slaying Two Dragons with One Stone – How to Prevent Cancer and Aging with the Same Strategy | AGING SCIENCES – Anti-Aging Firewalls
Pingback: Health through stressing fruits and vegetables – the Xenohormetic Live Food Hypothesis (updated) | AGING SCIENCES – Anti-Aging Firewalls