| The Part 1 blog entry discussed how TEs are activated by stresses, an initial step in the processes of genomic evolution. Stresses also activate alternative splicing.
The role of stress that I suggested above is supported by the 2015 publication Response of transposable elements to environmental stressors, which is concerned about stresses driving disease processes today. “Transposable elements (TEs) comprise a group of repetitive sequences that bring positive, negative, as well as neutral effects to the host organism. Earlier considered as “junk DNA,” TEs are now well-accepted driving forces of evolution and critical regulators of the expression of genetic information. — Evidence summarized in this review suggests that TEs are the sensitive endpoints for detection of effects caused by such environmental stressors, as ionizing radiation (terrestrial, space, and UV-radiation), air pollution (including particulate matter [PM]-derived and gaseous), persistent organic pollutants, and metals. Furthermore, the significance of these effects is characterized by their early appearance, persistence and presence in both, target organs and peripheral blood. Altogether, these findings suggest that TEs may potentially be introduced into safety and risk assessment and serve as biomarkers of exposure to environmental stressors. Furthermore, TEs also show significant potential to become invaluable surrogate biomarkers in clinic and possible targets for therapeutic modalities for disease treatment and prevention.”
Getting back to my views on evolution: The environmental conditions for every life form continues to evolve and change. Some change is fast, some slow. There are situations of relative stability, and life forms depend on these. But there can be no “forever” stability. Here again are some things I think about stress:
- Each life form at any time has adopted for survival in the semi-stable environment it exists in at that time. They calibrate their multiple internal homeostasis-seeking systems to function in that semi-stable environment. A thermoacidophile likes it very hot and acid. We can’t stand that. A methanogen likes it anaerobic with lots of methane, like in a sewer. We don’t. What we have in common is that we have DNA, RNA and have evolved according to some common mechanisms described here. Most everything else is different.
- Yet, because of constant flux, life forms experience a variety of stresses. Many of these will be mild and more or less expected; some are likely to be serious. Some will be wellness or life-threatening. A short draught or having insects eat a few if its leaves may stress a plant; having all of its leaves eaten by foraging deer may seriously threaten that plant; being overrun by a lava flow will kill it.
- Lifeforms have developed strategies for going on in the presences of change. Two important ones I focus on here are hormesis and evolution. Both of these strategies are triggered by stress.
Hormesis is the strategy used for surviving on the individual level, the capability of an organism or a biological subsystem to respond to a stress by mobilizing its defenses so as to actually improve its functionality, providing that the duration/intensity of the stress falls within a certain range. We have discussed hormesis extensively in this blog. In particular see 2012 Multifactorial hormesis, 2013 The Hormesis Bars, 2012 Radiation hormesis, 2012 Mitohormesis, and 2009 Hormesis and age retardation.
Evolution is a stress response mediated by transposable element and alternative splicing expression that leads to genomic change as discussed in this blog entry. This may effect individuals, germ lines, populations or species.
At least for certain common stress inducers (like heat, cold, hypoxia, oxidative load, pressure, presence of many toxins), the response of an organism is conditioned by the range of duration/intensity of the stressor
| Sub-hormetic stress |
little or no observable effect |
| Hormetic stress |
stress-response mechanisms of the organism provides healthy survival-giving response |
| TE-activating stress |
TEs are activated, leading to an enhanced probability of evolution on the genomic level |
| Lethal stress |
stress too great for organismic survival |
I do not know exactly how these ranges may overlap each other and I suspect that some stress responses are both hormetic and TE-activating. Impacts of a given stressor may be different in different organs and in different cells. And they will certainly be different in different individuals. And impacts may be determined by multiple variables such as initial redox state of the organism, circadian status, and other complicating factors. Remember the important distinction known to every engineering student but usually glossed over in the biomedical literature (and, I admit, glossed over here too), the difference between strain and stress.
- The external stimulus creating a challenge is called a strain, be this an insect attack on a plant a toxin ingested by your dog, or a fight with your boss.
- A stress is the response to the strain, and a given strain may be responded to very differently. Some plants of a given species may survive a given level of insect attack and some may die. A given dose of ingested castor oil may make some dogs of the same breed, weight and age only slightly sick and kill others. A fight with a boss may be shrugged off by one employee and lead to a mental breakdown in another. So stress is a function of system state as well as what is going on “our there.”
TEs: facilitators of evolution
Back while I was in 8th grade, I heard that evolution was due to a combination of random variation and natural selection for survival – whatever those meant. Later, perhaps early in college years, I learned further that the random variation was due to changes in DNA such as a DNA base pair mutation caused by a random oxidative event such as the passage of a cosmic ray. As time passed, it gradually became clear to me that there had to be lot more involved for evolution to happen. Evolution involves multiple complex and simultaneous adoptions and happens far more rapidly than can be explained by random molecule or gene-at-a-time events. Transposable elements in our genome – millions of different ones — provide a much more powerful framework for originating possibly useful mutations in our DNA, where long sequences can be exported within and across chromosomes. Sometimes a LINE-1 TE may pick up an unrelated segment of DNA and ship it “to who may be concerned.” Some times a gene may be duplicated that confers an evolutionary advantage, one like P53. TEs both provide a massive mechanism for introducing more than random variations, and they raise the question of the extent to which the “random variation” of evolution is really random, after all.
TEs can unquestionably mess up DNA, shipping large sequences into places where they seem to have no function other than to create problems. “LINE-1 expression damages host DNA via insertions and endonuclease-dependent DNA double-strand breaks (DSBs) that are highly toxic and mutagenic(ref).” Yet I side with some biologists who have been increasingly seeing TEs as facilitators of evolution.
The 2012 article Transposable Elements, Epigenetics, and Genome Evolution summarizes the situation. “Today, we know that TEs constitute more than half of the DNA in many higher eukaryotes. We know, too, that the fingerprints of TEs and transposition are everywhere in their genomes, from the coarsest features of genomic landscapes and how they change through real and evolutionary time to the finest details of gene structure and regulation. My purpose here is to challenge the current, somewhat pejorative, view of TEs as genomic parasites with the mounting evidence that TEs and transposition play a profoundly generative role in genome evolution. I contend that it is precisely the elaboration of epigenetic mechanisms from their prokaryotic origins as suppressors of genetic exchanges that underlies both the genome expansion and the proliferation of TEs characteristic of higher eukaryotes. This is the inverse of the prevailing view that epigenetic mechanisms evolved to control the disruptive potential of TEs. The evidence that TEs shape eukaryotic genomes is by now incontrovertible. My thesis, then, is that TEs and the transposases they encode underlie the evolvability of higher eukaryotes’ massive, messy genomes.” Epigenetic silencing of TEs is one of the topics Jim Watson treated in the Part 2 blog entry of this series.
Stresses lead to TE expression
Evolution appears to go far beyond protecting species. When the survival of a species begins to be seriously threatened by stresses, evolutionary innovation starts to happen. Evolutionary processes are kicked into gear which lead to multiple genetic variations in the species and possibly to the creation of new related species. Among such processes and perhaps key is the upgrading of expression of transposable elements. This has been observed in multiple species and with multiple kinds of stresses. Some examples are offered in these publications:
The 2012 publication Transposable elements: from DNA parasites to architects of metazoan evolution relates “The most unexpected insights that followed from the completion of the human genome a decade ago was that more than half of our DNA is derived from transposable elements (TEs). Due to advances in high throughput sequencing technologies it is now clear that TEs comprise the largest molecular class within most metazoan genomes. TEs, once categorised as “junk DNA”, are now known to influence genomic structure and function by increasing the coding and non-coding genetic repertoire of the host. In this way TEs are key elements that stimulate the evolution of metazoan genomes. This review highlights several lines of TE research including the horizontal transfer of TEs through host-parasite interactions, the vertical maintenance of TEs over long periods of evolutionary time, and the direct role that TEs have played in generating morphological novelty.”
The 2000 document Stress and transposable elements: co-evolution or useful parasites? relates: “In the natural world, individuals, populations and species all have to cope with environmental change. Individual organisms and their cells have to adapt physiologically through responses that are immediate and reversible. At the population and species levels, selection may lead to genetic changes and to the evolution of the inherited characteristics of an individual organism. In such organisms this long-term response is irreversible. — During the last two decades several authors have reported that stress increases the genetic variability of many quantitative traits in a population, see for instance Imasheva et al., (1998), including life history and morphology (Hoffmann & Parsons, 1997). This genetic variability may have various origins. For example, different genes may be expressed in normal as opposed to stressful environments. It has also been shown recently that genetic variability can be hidden by buffering proteins such as Hsp90 (Rutherford & Lindquist, 1998). This variability can be revealed by stress and then maintained when the protein function is restored. Finally, mutator mechanisms can be induced by stress. These are the origin of the genetic variability that allows selection to take place in response to environmental changes. At least two mechanisms are frequently described: those involving the SOS response (the activation of mutagenic activity) or the MRS response (inhibition of an antimutagenic system, the mismatch repair system) (Taddei et al., 1997) and those involving transposable elements (TEs) (Capy et al., 1997). Here we review and discuss findings of the impact of TEs on the host genome under stressful conditions and also summarize the various models put forward to account for these findings.”
1992 Induction of the mobile genetic element Dm-412 transpositions in the Drosophila genome by heat shock treatment “Males of a Drosophila melanogaster isogenic line with a mutation of the major gene for radius incompletus (ri) were treated by standard light heat shock (37 degrees C for 90 min) and by heavy heat shock (transfer of males from 37 degrees C for 2 hr to 4 degrees C for 1 hr and back; this procedure was repeated three times). In the F1 generation of treated males mated with nontreated females of the same isogenic line, mass transpositions of copia-like mobile genetic element Dm-412 were found. The altered positions of the element seem nonrandom; five “hot spots” of transposition were found. Probabilities of transpositions were estimated after light heat shock and heavy heat shock and in the control sample. These probabilities were, respectively, 3.4 x 10(-2), 8.7 x 10(-2), and less than 4.1 x 10(-4) transpositions per genome per occupied position per generation. Therefore, as a result of heat shock treatment, the probabilities of transpositions were two orders of magnitude greater than those of the control sample in the next generation after induction. Comparison of the results with those after stepwise temperature treatment shows that the induction depends on the intensity of the stress action (temperature treatment) rather than on the type of the stress action.”
2013 Transposable elements: powerful contributors to angiosperm evolution and diversity. “Transposable elements (TEs) are a dominant feature of most flowering plant genomes. Together with other accepted facilitators of evolution, accumulating data indicate that TEs can explain much about their rapid evolution and diversification. Genome size in angiosperms is highly correlated with TE content and the overwhelming bulk (>80%) of large genomes can be composed of TEs. Among retro-TEs, long terminal repeats (LTRs) are abundant, whereas DNA-TEs, which are often less abundant than retro-TEs, are more active. Much adaptive or evolutionary potential in angiosperms is due to the activity of TEs (active TE-Thrust), resulting in an extraordinary array of genetic changes, including gene modifications, duplications, altered expression patterns, and exaptation to create novel genes, with occasional gene disruption. TEs implicated in the earliest origins of the angiosperms include the exapted Mustang, Sleeper, and Fhy3/Far1 gene families. Passive TE-Thrust can create a high degree of adaptive or evolutionary potential by engendering ectopic recombination events resulting in deletions, duplications, and karyotypic changes. TE activity can also alter epigenetic patterning, including that governing endosperm development, thus promoting reproductive isolation. Continuing evolution of long-lived resprouter angiosperms, together with genetic variation in their multiple meristems, indicates that TEs can facilitate somatic evolution in addition to germ line evolution. Critical to their success, angiosperms have a high frequency of polyploidy and hybridization, with resultant increased TE activity and introgression, and beneficial gene duplication. Together with traditional explanations, the enhanced genomic plasticity facilitated by TE-Thrust, suggests a more complete and satisfactory explanation for Darwin’s “abominable mystery”: the spectacular success of the angiosperms.”
Rapid mobilization of TEs by stresses may be critical for the success of invasive species.
2015 Transposable elements as agents of rapid adaptation may explain the genetic paradox of invasive species “Rapid adaptation of invasive species to novel habitats has puzzled evolutionary biologists for decades, especially as this often occurs in the face of limited genetic variability. Although some ecological traits common to invasive species have been identified, little is known about the possible genomic/genetic mechanisms that may underlie their success. A common scenario in many introductions is that small founder population sizes will often lead to reduced genetic diversity, but that invading populations experience large environmental perturbations, such as changes in habitat and environmental stress. Although sudden and intense stress is usually considered in a negative context, these perturbations may actually facilitate rapid adaptation by affecting genome structure, organization and function via interactions with transposable elements (TEs), especially in populations with low genetic diversity. Stress-induced changes in TE activity can alter gene action and can promote structural variation that may facilitate the rapid adaptation observed in new environments. We focus here on the adaptive potential of TEs in relation to invasive species and highlight their role as powerful mutational forces that can rapidly create genetic diversity. We hypothesize that activity of transposable elements can explain rapid adaptation despite low genetic variation (the genetic paradox of invasive species) — .”
2014 Transposable element islands facilitate adaptation to novel environments in an invasive species. This one is about an ant species.
Stress-activated TEs and the other pro-evolutionary mechanisms characterized in this blog entry also play roles in the stress responses of individuals for maintaining homeostasis and in disease processes. These topics are not just about what went on during tens of millions of years of evolution, They are also about what goes on in us right now.
An example of this is reported in the 2015 publication Stress and the dynamic genome: Steroids, epigenetics, and the transposome: “Stress plays a substantial role in shaping behavior and brain function, often with lasting effects. How these lasting effects occur in the context of a fixed postmitotic neuronal genome has been an enduring question for the field. Synaptic plasticity and neurogenesis have provided some of the answers to this question, and more recently epigenetic mechanisms have come to the fore. The exploration of epigenetic mechanisms recently led us to discover that a single acute stress can regulate the expression of retrotransposons in the rat hippocampus via an epigenetic mechanism. We propose that this response may represent a genomic stress response aimed at maintaining genomic and transcriptional stability in vulnerable brain regions such as the hippocampus. This finding and those of other researchers have made clear that retrotransposons and the genomic plasticity they permit play a significant role in brain function during stress and disease. These observations also raise the possibility that the transposome might have adaptive functions at the level of both evolution and the individual organism.”
In plants as well as animals, stresses activate TEs wich results in higher expression of stress responsive genes. For example, the1997 document The expression of the tobacco Tnt1 retrotransposon is linked to plant defense responses reports: “Activation of retrotransposons by stresses and external changes is common in all eukaryotic systems, including plants. The transcription of the tobacco Tnt1 retrotransposon was studied in its natural host as well as in Arabidopsis and tomato. It is activated by factors of microbial origin, by external stresses, and by viral, bacterial, and fungal attacks. Tnt1 expression is linked with the biological responses of the plant to the elicitor or to the pathogen attack and in particular with the early steps of the metabolic pathways leading to the activation of plant defense genes. In most cases, the basic features of Tnt1 regulation in tobacco are maintained in tomato and Arabidopsis, but some host-specific regulations were shown.”
2013 How do mammalian transposons induce genetic variation? A conceptual framework: the age, structure, allele frequency, and genome context of transposable elements may define their wide-ranging biological impacts. “We present a conceptual framework to understand how the ages, allele frequencies, molecular structures, and especially the genomic context of mammalian TEs each can influence their various possible functional consequences. While most TEs are ancient relics, certain classes can move from one chromosomal location to another even now. Indeed, striking recent data show that extensive transposition occurs not only in the germline over evolutionary time, but also in developing somatic tissues and particular human cancers. While occasional germline TE insertions may contribute to genetic variation, many other, similar TEs appear to have little or no impact on neighboring genes. However, the effects of somatic insertions on gene expression and function remain almost completely unknown.”
Additional references:
2009 The impact of retrotransposons on human genome evolution
2012 Presidential address. Transposable elements, epigenetics, and genome evolution.
The evolution of epigenetics.
1995 LTR-retrotransposons and MITEs: important players in the evolution of plant genomes.
2013 Abundance and distribution of transposable elements in two Drosophila QTL mapping resources.
But TEs are only part of the picture as we will see;
Other key biological mechanisms work with TEs to support evolution (and normal biological functioning)
As pointed out above, the important role that TEs seem to play in evolution is that of selectively enhancing genetic diversity in response to stress. This begs the question of exactly how that genetic diversity can lead to enhanced survivability. What are the evolutionarily conserved mechanisms of “survival of the fittest?” Is genomic evolution purely a matter of trial and error once TEs mix-and-match and spread the DNA around?
No. Absolutely not. We could ever have gotten this far with evolution if that were the case, even after 450 million years. A picture of the situation is beginning to emerge, but it carries us off into frontier areas of genomic research. The answers are far from being all in but we already have quite a bit to work with. Among the mechanisms that work with TEs are other processes that go on in DNA, including incRNAs, alternative splicing. A to I editing, and exonization. I briefly characterize these and then go on to illustrate how they contribute to evolution along with TEs by quoting from a variety of research publications
Long non-coding RNAs – incRNAs and lincRNAs
incRNAs are long non-coding RNAs generally greater than 200 nucleotides in length, an important species in the RNA Zoo known for their regulatory functions and possible association with cellular senescence(ref)(ref). lincRNAs are long intergenic noncoding RNAs(ref). That is, they live in the regions of DNA between genes.
Recent research suggests that TE insertions contribute significantly to the DNA found in long non-coding RNAs. This can explain the rapid gene evolution observed in long non-coding RNAs.
A very recent chapter of the TE-incRNA-evolution story has to do with SINEUPs, as told by this 2015 publication SINEUPs: A new class of natural and synthetic antisense long non-coding RNAs that activate translation. “Over the past 10 years, it has emerged that pervasive transcription in mammalian genomes has a tremendous impact on several biological functions. Most of transcribed RNAs are lncRNAs and repetitive elements. In this review, we will detail the discovery of a new functional class of natural and synthetic antisense lncRNAs that stimulate translation of sense mRNAs. These molecules have been named SINEUPs since their function requires the activity of an embedded inverted SINEB2 sequence to UP-regulate translation. Natural SINEUPs suggest that embedded Transposable Elements may represent functional domains in long non-coding RNAs. Synthetic SINEUPs may be designed by targeting the antisense sequence to the mRNA of choice representing the first scalable tool to increase protein synthesis of potentially any gene of interest. We will discuss potential applications of SINEUP technology in the field of molecular biology experiments, in protein manufacturing as well as in therapy of haploinsufficiencies.”
The theme in this publication of TEs contributing significant insertions into long non-coding RNAs is also supported by this 2015 publication: Transposable Element Insertions in Long Intergenic Non-Coding RNA Genes. “Transposable elements (TEs) are abundant in mammalian genomes and appear to have contributed to the evolution of their hosts by providing novel regulatory or coding sequences. We analyzed different regions of long intergenic non-coding RNA (lincRNA) genes in human and mouse genomes to systematically assess the potential contribution of TEs to the evolution of the structure and regulation of expression of lincRNA genes. Introns of lincRNA genes contain the highest percentage of TE-derived sequences (TES), followed by exons and then promoter regions although the density of TEs is not significantly different between exons and promoters. Higher frequencies of ancient TEs in promoters and exons compared to introns implies that many lincRNA genes emerged before the split of primates and rodents. The content of TES in lincRNA genes is substantially higher than that in protein-coding genes, especially in exons and promoter regions. A significant positive correlation was detected between the content of TEs and evolutionary rate of lincRNAs indicating that inserted TEs are preferentially fixed in fast-evolving lincRNA genes. These results are consistent with the repeat insertion domains of LncRNAs hypothesis under which TEs have substantially contributed to the origin, evolution, and, in particular, fast functional diversification, of lincRNA genes.”
The same theme is articulated in the 2014 publication The RIDL hypothesis: transposable elements as functional domains of long noncoding RNAs. “Our genome contains tens of thousands of long noncoding RNAs (lncRNAs), many of which are likely to have genetic regulatory functions. It has been proposed that lncRNA are organized into combinations of discrete functional domains, but the nature of these and their identification remain elusive. One class of sequence elements that is enriched in lncRNA is represented by transposable elements (TEs), repetitive mobile genetic sequences that have contributed widely to genome evolution through a process termed exaptation. Here, we link these two concepts by proposing that exonic TEs act as RNA domains that are essential for lncRNA function. We term such elements Repeat Insertion Domains of LncRNAs (RIDLs). A growing number of RIDLs have been experimentally defined, where TE-derived fragments of lncRNA act as RNA-, DNA-, and protein-binding domains. We propose that these reflect a more general phenomenon of exaptation during lncRNA evolution, where inserted TE sequences are repurposed as recognition sites for both protein and nucleic acids. We discuss a series of genomic screens that may be used in the future to systematically discover RIDLs. The RIDL hypothesis has the potential to explain how functional evolution can keep pace with the rapid gene evolution observed in lncRNA. More practically, TE maps may in the future be used to predict lncRNA function.”
So, TEs can insert segments in incRNAs. What are the downstream consequences of this in terms of incRNA regulation of transcription and ultimately evolution? This diagram illustrates various ways in which incRNAs can regulate transcription:
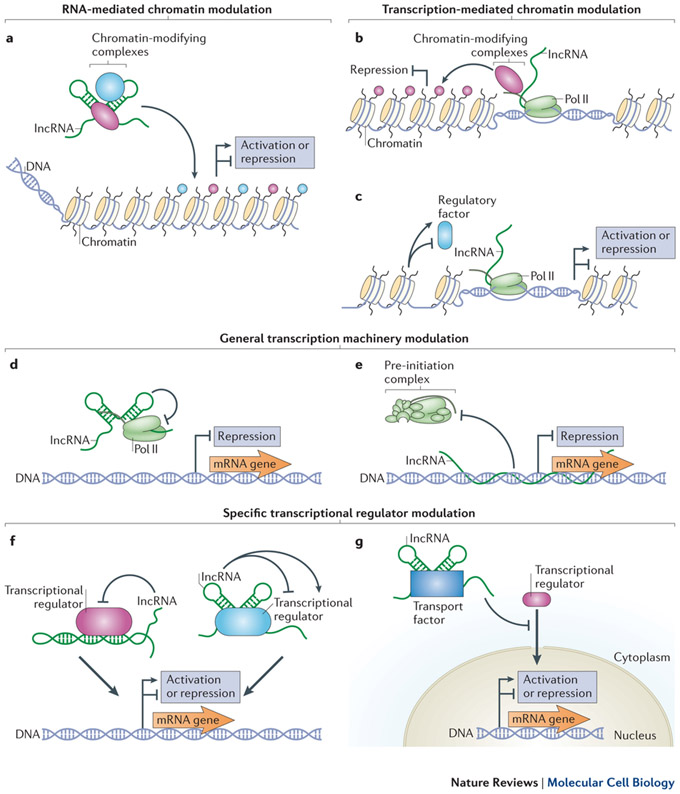
Image and legend source: 2013 RNA in unexpected places: long non-coding RNA functions in diverse cellular contexts. “ a–c | Long non-coding RNAs (lncRNAs) can modulate chromatin through transcription-independent (part a) and transcription-dependent mechanisms (parts b and c). lncRNAs can bind one or more chromatin-modifying complexes and target their activities to specific DNA loci (part a). Depending on the nature of the enzymes bound, lncRNA-mediated chromatin modifications can activate or repress gene expression22, 23, 26, 27, 120. Chromatin-modifying complexes bound to the RNA polymerase II (Pol II) carboxy-terminal domain (CTD) can modify chromatin during transcription of lncRNAs33, 34, 35 (part b). Transcription of lncRNAs can also result in chromatin remodelling that can either favour or inhibit the binding of regulatory factors (part c). Depending on the nature of the factors that bind during remodelling, gene expression is activated or repressed37, 38, 39, 40. d–g | lncRNAs can modulate both the general transcription machinery (parts d and e) as well as specific regulatory factors (parts f and g). lncRNAs can bind Pol II directly to inhibit transcription47 (part d). Formation of lncRNA–DNA triplex structures can also inhibit the assembly of the pre-initiation complex48 (part e). lncRNAs can fold into structures that mimic DNA-binding sites (left) or that generally inhibit or enhance the activity of specific transcription factors (right)50, 51, 52, 53 (part f). lncRNAs can also regulate gene expression by binding specific transport factors to inhibit the nuclear localization of specific transcription factors54 (part g).”
Alternative splicing
Up through the mid-late 70s, biologists thought that a single gene could make only one unique protein. Now we know that a single gene can make tens or hundreds of thousands of different proteins, The secret is alternative splicing together of gene components – exons or introns converted to exons – going into the translation phase of protein making. Alternative splicing is not “alternative.” It is the regular way biology operates.
“Alternative splicing is a regulated process during gene expression that results in a single gene coding for multiple proteins. In this process, particular exons of a gene may be included within or excluded from the final, processed messenger RNA (mRNA) produced from that gene.[1] Consequently the proteins translated from alternatively spliced mRNAs will contain differences in their amino acid sequence and, often, in their biological functions (see Figure). Notably, alternative splicing allows the human genome to direct the synthesis of many more proteins than would be expected from its 20,000 protein-coding genes. Alternative splicing is sometimes termed differential splicing. — Alternative splicing occurs as a normal phenomenon in eukaryotes, where it greatly increases the biodiversity of proteins that can be encoded by the genome;[1] in humans, ~95% of multi-exonic genes are alternatively spliced.[2] There are numerous modes of alternative splicing observed, of which the most common is exon skipping. In this mode, a particular exon may be included in mRNAs under some conditions or in particular tissues, and omitted from the mRNA in others.[1]”
Quoted text and illustration from Wikipedia
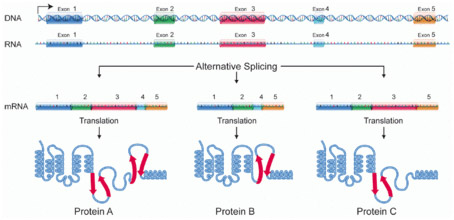
Alternative splicing produces three protein isoforms.
Alternative splicing is evolutionarily conserved, known to be applicable in insects as well as in mammals including us(ref). Although the mechanisms of it have become more sophisticated in higher lifeforms, alternative splicing is ancient and possibly came into being as an essential feature of multi-celled organisms. The 2004 the publication How did alternative splicing evolve? relates: “Alternative splicing creates transcriptome diversification, possibly leading to speciation. A large fraction of the protein-coding genes of multicellular organisms are alternatively spliced, although no regulated splicing has been detected in unicellular eukaryotes such as yeasts. A comparative analysis of unicellular and multicellular eukaryotic 5′ splice sites has revealed important differences — the plasticity of the 5′ splice sites of multicellular eukaryotes means that these sites can be used in both constitutive and alternative splicing, and for the regulation of the inclusion/skipping ratio in alternative splicing. So, alternative splicing might have originated as a result of relaxation of the 5′ splice site recognition in organisms that originally could support only constitutive splicing.”
Alternative splicing is extremely important, not only for evolution but also for health and longevity. James Watson and I are currently working on blog entries on it which I expect we will publish shortly.
A to I editing
Adenosine-to-inosine (A-to-I) RNA editing is the most important form of RNA editing in us. It is not an oddball phenomenon, but has played an important role in making us human. The 2014 publication A-to-I RNA editing occurs at over a hundred million genomic sites, located in a majority of human genes describes the mechanism and its importance: “RNA molecules transmit the information encoded in the genome and generally reflect its content. Adenosine-to-inosine (A-to-I) RNA editing by ADAR proteins converts a genomically encoded adenosine into inosine. It is known that most RNA editing in human takes place in the primate-specific Alu sequences, but the extent of this phenomenon and its effect on transcriptome diversity are not yet clear. Here, we analyzed large-scale RNA-seq data and detected ∼1.6 million editing sites. As detection sensitivity increases with sequencing coverage, we performed ultradeep sequencing of selected Alu sequences and showed that the scope of editing is much larger than anticipated. We found that virtually all adenosines within Alu repeats that form double-stranded RNA undergo A-to-I editing, although most sites exhibit editing at only low levels (<1%). Moreover, using high coverage sequencing, we observed editing of transcripts resulting from residual antisense expression, doubling the number of edited sites in the human genome. Based on bioinformatic analyses and deep targeted sequencing, we estimate that there are over 100 million human Alu RNA editing sites, located in the majority of human genes. These findings set the stage for exploring how this primate-specific massive diversification of the transcriptome is utilized.”
Here is an illustration of A to I editing.
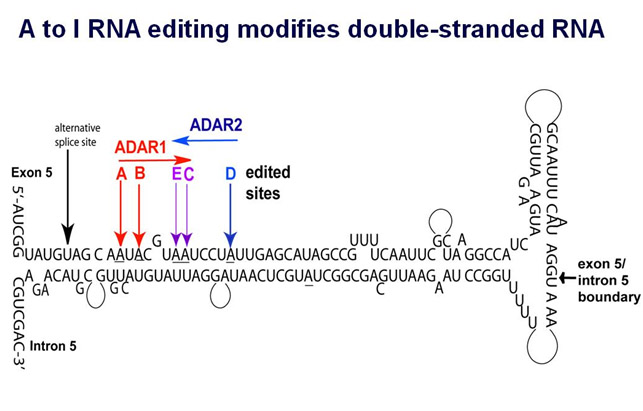
Image source “ The double-stranded RNA loop of the human 5-HT2C receptor transcript. ADAR enzymes have been shown to bind only to double-stranded RNA. The specific sites of activity of ADARs 1 and 2 are indicated.”
Note that we have discussed Alu TE elements in the Part2 blog entry
“Conversion of a genetically encoded adenosine (A) into an inosine (I) while preserving sequence in RNA is accomplished by adenosine deaminases (ADARs). “The primate specific Alu sequences are the dominant short interspersed nuclear element (SINEs) in the primate genomes (International Human Genome Sequencing Consortium 2001; Cordaux and Batzer 2009). Humans have about a million copies of Alu, roughly 300 bp long each, accounting for ∼10% of their genome. Since these repeats are so common, especially in gene-rich regions (Korenberg and Rykowski 1988), pairing of two oppositely oriented Alus located in the same pre-mRNA structure is likely. Such pairing produces a long and stable dsRNA structure, an ideal target for the ADARs. Indeed, recent studies have shown that Alu repeats account for >99% of editing events found so far in humans (Athanasiadis et al. 2004; Blow et al. 2004; Kim et al. 2004; Levanon et al. 2004; Ramaswami et al. 2012, 2013).”
This 2008 publication RNA editing in regulating gene expression in the brain discusses how A to I editing and alternative splicing have had ” profound importance for normal nervous system function in a wide range of invertebrate and vertebrate model organisms.”
Exonization
“A funny thing happened to my RNA on the way to making a protein”
Exonization can convert an intron in a segment of RNA into an exon, which means that instead of being left out during translation it gets converted into a protein-encoding segment of DNA. If the intron so-converted comes from a TE insertion, an alternative splicing and possible new protein results. The result can frequently be negative.
The 2011 publication Exonization of transposed elements: A challenge and opportunity for evolution relates: “Protein-coding genes are composed of exons and introns flanked by untranslated regions. Before the mRNA of a gene can be translated into protein, the splicing machinery removes all the intronic regions and joins the protein-coding exons together. Exonization is a process, whereby genes acquire new exons from non-protein-coding, primarily intronic, DNA sequences. Genomic insertions or point mutations within DNA sequences often generate alternative splice sites, causing the splicing system to include new sequences as exons or to elongate existing exons. Because the alternative splice sites are not as efficient as the originals the new variants usually constitute a minor fraction of mature mRNAs. While the prevailing original splice variant maintains functionality, the additional sequence, free from selection pressure, evolves a new function or eventually vanishes. If the new splice variant is advantageous, selection might operate to optimize the new splice sites and consequently increase the proportion of the alternative splice variant. In some instances, the original splice variant is completely replaced by constitutive splicing of the new form. Because of the fortuitous presence of internal splice site-like structures within their sequences, portions of transposed elements frequently serve as modules of exonization. Their recruitment requires a long and versatile optimization process involving multiple changes over a time span of millions, even hundreds of millions, of years. Comparisons of corresponding genes and mRNAs in phylogenetically related species enables one to chronologically reconstruct such changes, from ancient ancestors to living species, in a stepwise manner. We will review this process using three different exemplary cases: (1) the evolution of a constitutively spliced mammalian-wide repeat (MIR), (2) the evolution of an alternative exon 1 from an alternative 5′-extended primary transcript containing an Alu element, and (3) a rare case of the stepwise exoniztion of an Alu element-derived sequence mediated by A-to-I RNA editing.”
The 2011 publication Characteristics of transposable element exonization within human and mouse reported “Insertion of transposed elements within mammalian genes is thought to be an important contributor to mammalian evolution and speciation. Insertion of transposed elements into introns can lead to their activation as alternatively spliced cassette exons, an event called exonization. Elucidation of the evolutionary constraints that have shaped fixation of transposed elements within human and mouse protein coding genes and subsequent exonization is important for understanding of how the exonization process has affected transcriptome and proteome complexities. Here we show that exonization of transposed elements is biased towards the beginning of the coding sequence in both human and mouse genes. Analysis of single nucleotide polymorphisms (SNPs) revealed that exonization of transposed elements can be population-specific, implying that exonizations may enhance divergence and lead to speciation. SNP density analysis revealed differences between Alu and other transposed elements. Finally, we identified cases of primate-specific Alu elements that depend on RNA editing for their exonization. These results shed light on TE fixation and the exonization process within human and mouse genes.”
Some supporting research publications
Italics in quoted segments of text are my own, for emphasis of particularly important points.
TEs contribute to evolution by furnishing functional intron and exon domains to long non-coding RNAs (incRNAs)
TEs, A to I editing, alternative splicing, exonization and evolution
How do the DNA processes of A to I editing, alternative splicing, exonization together with TE expression come together to support the sweeping view of evolution that I outlined in the introduction to this blog – that our genome consists of a gigantic archive of mostly-unused DNA segments from our evolutionary history that can be repurposed and recombined in the interest of evolution? The following publications suggest how, each with their own viewpoints:
The 2015 publication Alternative splicing and co-option of transposable elements: the case of TMPO/LAP2α and ZNF451 in mammals relates: “Transposable elements constitute a large fraction of vertebrate genomes and, during evolution, may be co-opted for new functions. Exonization of transposable elements inserted within or close to host genes is one possible way to generate new genes, and alternative splicing of the new exons may represent an intermediate step in this process. The genes TMPO and ZNF451 are present in all vertebrate lineages. Although they are not evolutionarily related, mammalian TMPO and ZNF451 do have something in common-they both code for splice isoforms that contain LAP2alpha domains. We found that these LAP2alpha domains have sequence similarity to repetitive sequences in non-mammalian genomes, which are in turn related to the first ORF from a DIRS1-like retrotransposon. This retrotransposon domestication happened separately and resulted in proteins that combine retrotransposon and host protein domains. The alternative splicing of the retrotransposed sequence allowed the production of both the new and the untouched original isoforms, which may have contributed to the success of the colonization.”
There we have it. Combine TE insertions, exonization and alternative splicing, and it seems we can come up with new genes. The following documents tell important additional parts of the story about how we made it to becoming humans.
Remember what happened to your ancestors 40-60 million years ago? The 2008 publication Beyond DNA: RNA editing and steps toward Alu exonization in primates discusses how Alu TE elements, A to I editing and alternative splicing worked together to create the gorilla, the chimp, many other of our primate cousins, and us. “The exaptation of transposed elements into protein-coding domains by a process called exonization is one important evolutionary pathway for generating novel variant functions of gene products. Adenosine-to-inosine (A-to-I) modification is a recently discovered, RNA-editing-mediated mechanism that contributes to the exonization of previously unprocessed mRNA introns. In the human nuclear prelamin A recognition factor gene transcript, the alternatively spliced exon 8 results from an A-to-I editing-generated 3′ splice site located within an intronic Alu short interspersed element. Sequence comparisons of representatives of all primate infraorders revealed the critical evolutionary steps leading to this editing-mediated exonization. The source of exon 8 was seeded within the primary transcript about 58-40 million years ago by the head-to-head insertions of two primate-specific Alu short interspersed elements in the common ancestor of anthropoids. The latent protein-coding potential was realized 34-52 million years later in a common ancestor of gorilla, chimpanzee, and human as a result of numerous changes at the RNA and DNA level. Comparisons of 426 processed mRNA clones from various primate species with their genomic sequences identified seven different RNA-editing-mediated alternative splice variants. In total, 30 A-to-I editing sites were identified. The gorilla, chimpanzee, and human nuclear prelamin A recognition factor genes exemplify the versatile interplay of pre- and posttranscriptional modifications leading to novel genetic potential.”
Another part of the same story is told in the 2013 publication related to TE DNA insertions, long non-coding RNAs, simian and our evolution is ANRIL/CDKN2B-AS shows two-stage clade-specific evolution and becomes conserved after transposon insertions in simians. “BACKGROUND: Many long non-coding RNA (lncRNA) genes identified in mammals have multiple exons and functional domains, allowing them to bind to polycomb proteins, DNA methyltransferases, and specific DNA sequences to regulate genome methylation. Little is known about the origin and evolution of lncRNAs. ANRIL/CDKN2B-AS consists of 19 exons on human chromosome 9p21 and regulates the expression of three cyclin-dependent kinase inhibitors (CDKN2A/ARF/CDKN2B). — RESULTS:ANRIL/CDKN2B-AS originated in placental mammals, obtained additional exons during mammalian evolution but gradually lost them during rodent evolution, and reached 19 exons only in simians. ANRIL lacks splicing signals in mammals. In simians, multiple transposons were inserted and transformed into exons of the ANRIL gene, after which ANRIL became highly conserved. A further survey reveals that multiple transposons exist in many lncRNAs. — CONCLUSIONS:ANRIL shows a two-stage, clade-specific evolutionary process and is fully developed only in simians. The domestication of multiple transposons indicates an impressive pattern of “evolutionary tinkering” and is likely to be important for ANRIL’s structure and function. The evolution of lncRNAs and that of transposons may be highly co-opted in primates. Many lncRNAs may be functional only in simians.” We became smart and capable monkeys, poised later to evolve further into us humans.
You can also check out the publication One hundred million adenosine-to-inosine RNA editing sites: hearing through the noise. “A small subset of edited Alu elements has been shown to exhibit diverse functional roles in the regulation of alternative splicing, miRNA repression, and cis-regulation of distant RNA editing sites. The low level of editing for the remaining majority may be non-functional, yet their persistence in the primate genome provides enhanced genomic flexibility that may be required for adaptive evolution.”
The evolution-promoting actions of the mechanisms described here work in plants too. The 2012 publication Genome-wide survey of ds exonization to enrich transcriptomes and proteomes in plants relates “Insertion of transposable elements (TEs) into introns can lead to their activation as alternatively spliced cassette exons, an event called exonization which can enrich the complexity of transcriptomes and proteomes. — The insertion patterns of Ds and the polymorphic splice donor sites increased the transcripts and subsequent protein isoforms. Protein isoforms contain protein sequence due to unspliced intron-TE region and/or a shift of the reading frame. The number of interior protein isoforms would be twice that of C-terminal isoforms, on average. TE exonization provides a promising way for functional expansion of the plant proteome.”
There are many additional relevant citations such as:
2012 Intronic retroelements: Not just “speed bumps” for RNA polymerase II.
2013 Retroelements in human disease.
2005 Gene-breaking: a new paradigm for human retrotransposon-mediated gene evolution.
2011 Intronic L1 retrotransposons and nested genes cause transcriptional interference by inducing intron retention, exonization and cryptic polyadenylation.
2009 Exon-trapping mediated by the human retrotransposon SVA.
2007 Biased exonization of transposed elements in duplicated genes: A lesson from the TIF-IA gene.
2010 Transposable elements in disease-associated cryptic exons.
2009 Disease-causing mutations improving the branch site and polypyrimidine tract: pseudoexon activation of LINE-2 and antisense Alu lacking the poly(T)-tail.
2006 Identification of multiple transcription initiation, polyadenylation, and splice sites in the Drosophila melanogaster TART family of telomeric retrotransposons.
2004 Activation of cryptic 3′ splice sites within introns of cellular genes following gene entrapment.
2010 Functions and regulation of RNA editing by ADAR deaminases.
2010 Genomic gems: SINE RNAs regulate mRNA production.
2009 Transcription of the rat testis-specific Rtdpoz-T1 and -T2 retrogenes during embryo development: co-transcription and frequent exonisation of transposable element sequences.
I expect that we will discuss several of the intriguing ideas raised in these publications in forthcoming blog entries
On the way to a Grand Unified Theory of biology and aging
In November 2013, I published a blog entry treatise Prospectus for a Grand Unified theory of Biology, Health and Aging. The discussion was focused on the possibility of identifying unifying principles of biology that provide a simple basic structure for this field of incredible complexity. The concept of a GUTb was introduced earlier in my PowerPoint presentation Multifactorial Hormesis which examines the roles of stresses and stress responses in biology.
The closest thing to a GUTb we have ever had in biology is Darwin’s theory of evolution, and it is precisely because of the importance of TEs to evolution that I was originally motivated to write this blog entry. To start, I list some properties I would like a theory to have to qualify as a GUT. A Grand Unified Theory of Biology (GUTb) must be concerned with universals.
- It should apply to all life forms ranging from primitive viruses, molds and bacteria to plants, ants, gnats, mice. lice, whales, snails, dogs, frogs, hogs, monkeys, donkeys and all other animals including us. And methanogens, certainhalophiles and thermoacidophiles and lot of other entities with strange names. That is, it should apply to all entities in the three main categories of life: bacteria, archaea and the eukaryotes, which include animals, plants and fungi.
- It should apply to all levels of biological organization (e.g. molecular, cell, organ, system, whole organism and organism in social context).
- It should be interesting, exciting, have predictive power and be consistent with all we know, including about evolution. It should apply historically, for now, and for the foreseeable future.
- It should provide us with insights into what all life is about and help guide our research.
I believe what is said in this blog entry related to TEs, non-coding RNAs, alternative splicing, A to I editing, exonization and evolution stands up to these criteria.
In the process of researching these blogs, I believe we are beginning to discern the basic mechanisms of evolution – what kicks it, off, the genomic steps of it happening, and even more fundamental – what it is: how it works in species, in populations and even within individuals. I believe Jim Watson joins me in finding this very exciting. I think a GUT of biology is no more about specific processes in specific organelles, cells or organisms – rules, exceptions, and exceptions to exceptions – than a GUT of physics is about the specific ways specific substances move when subject to specific forces. We can catalog, analyze and describe to our heart’s desire but this will always be about specific organisms living in particular circumstances at particular historical periods – the endpoint examples of evolution, not the process of how it works. Rather the GUT should be about how evolution itself takes place, about how new biological entities, rules and even species can come into being when needed. And about how they can go out of being when no longer needed, about mosaicism, about keeping an accessible record of what happened since the start of it all, about repurposing of solutions, and about the insatiable drive of biological entities not only for survival but also for improvement.
I don’t want to come across as saying I know a lot about evolution. I don’t. What I do want to convey is that I experience a lot of excitement about what I am learning discussed here.
We expect our readers will discover that these blog entries are not just about what happened 150 and 25 million years ago, and glass jars in musty museums with specimens in them. They are about little-discussed but extremely important mechanisms that affect our health and longevity right now, and that point to new interventions beyond those discussed in the usual medical or longevity literature. We plan to discuss specific health and longevity implications of alternative splicing in the following two blog entries.
These new blog entries will be centered on the same biological processes discussed here and in the previous blog entries in this series, alternative splicing, TEs, incRNAs, A to I editing and exonization – but focused on disease processes and human health instead of on evolution. Jim Watson and I think that alternative splicing plays a major role in determining health span and lifespan. We discuss how alternative splicing controls protein diversity, protein localization, protein function, and aging. And we discuss how ubiquitous alternative splicing is, playing key roles in such diverse areas as gender determination and autism, and in several key longevity-related pathways. Alternative splicing is a critical feature of both the IGF-1 gene, the high affinity IGF-2 protein found in cancer, and the IGF-1 receptor (and the insulin receptor). Alternative splicing may explain much of the difference between super centenarians and “regular people” who do not carry longevity-related mutations, It could account for the differences in longevity in those of us who do not have a heterozygous “Loss-of-function” mutation in our IGF-1R gene. The next blog entry in particular, written by Jim Watson, will focus on Hutchinson-Gilford Progeria Syndrome (HGPS), a model of aging where children go through all the steps of aging and die as old men and women from diseases of old age before their 20th birthdays. The key matter involved is hereditary alternative splicing of a single gene, Lamin A. |





Pingback: 2017 meeting of the International Dose-Response Society | AGINGSCIENCES™ – Anti-Aging Firewalls™
Pingback: SHIT DOESN’T JUST HAPPEN – ON CREATION, NATURAL AND HUMAN – BEING AND CREATION BLOG
Pingback: ON AGING - AGINGSCIENCES™ - Anti-Aging Firewalls™AGINGSCIENCES™ – Anti-Aging Firewalls™